Scientists Awarded $6.6 Million for Research in Computational Biology
Three groups at The Scripps Research Institute (TSRI) have been awarded grants from the National Institutes of Health (NIH) to develop methods for computational modeling and to apply them to cutting-edge systems in biology and health.
“The three projects are highly symbiotic, each addressing a different state-of-the-art challenge in computational biology, but built using a common computational framework that will allow facile collaboration between the groups,” said Professor Arthur Olson, founder of the Molecular Graphics Laboratory, which is currently part of the TSRI Department of Integrative Structural and Computational Biology.
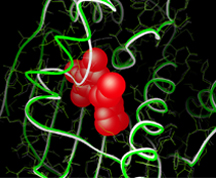
Stefano Forli, assistant professor of integrative structural and computational biology, was awarded $2.7 million to continue development of AutoDock, the most widely used method for computational docking of drugs and inhibitors to targets of medicinal interest. The proposed work will include improvements to the scoring function, allowing faster and more accurate prediction of how drugs act, new tools for advanced structure-based design of drugs and simplified methods for use by a wide community of non-expert users.
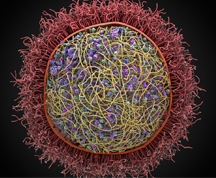
David S. Goodsell, associate professor of molecular biology, was awarded $2.3 million to develop new methods for modeling the molecular structure of entire cells. The size and complexity of these structural models is unprecedented, and the project is currently leveraging methods developed by the gaming community. In collaboration with experimental scientists, the methods will be used to study bacterial nucleoid structure, cell division and other cellular processes.
Michel F. Sanner, associate professor of molecular biology, initiated a research project addressing a significant challenge: the incorporation of the dynamic nature of proteins into docking simulations. Years six through nine of this project have been funded with an award of $1.6 million. His group will continue development of AutoDockFR, a fast computational docking method with advanced features, to represent local and global receptor motions during binding of flexible drug-like molecules to biomolecular targets.
Send comments to: press[at]scripps.edu