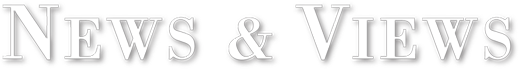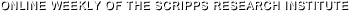Team Reveals How Deadly Ebola Virus Assembles
Findings Challenge Conventional Wisdom and Offer Multiple Drug Targets
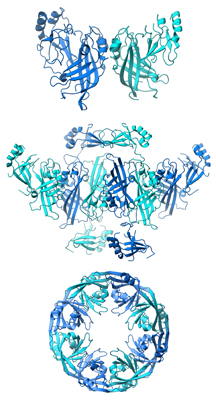
Scientists at The Scripps Research Institute (TSRI) have discovered the molecular mechanism by which the deadly Ebola virus assembles, providing potential new drug targets. Surprisingly, the study showed that the same molecule that assembles and releases new viruses also rearranges itself into different shapes, with each shape controlling a different step of the virus’s life cycle.
“Like a ‘Transformer’, this protein of the Ebola virus adopts different shapes for different functions,” said Erica Ollmann Saphire, professor in the Department of Immunology and Microbial Science at TSRI. “It revises a central dogma of molecular biology—that a protein molecule has one shape that predestines one biological function."
The research was published today in the peer-reviewed journal Cell.
“These findings open doors to developing new drugs against Ebola,” added Zachary Bornholdt, senior staff scientist and first author of the study. “Drugs to block viral replication could target any of the structures themselves or the intermediate steps in the structural transformation process.”
Ebola hemorrhagic fever is one of the most virulent diseases known to humankind. Very few pathogens prove more dangerous than Ebola virus once a person is infected. There is no cure, and the case-fatality rate can be up to 90 percent, depending on which strain is involved.
Ebola virus and its cousin Marburg virus are spread when people come into contact with the bodily fluids of a person or animal who is already infected. Infection causes rapidly progressing high fever, hemorrhage and shock. No drugs or vaccines are yet available for human use. Currently, the standard treatment consists of administering fluids and taking protective measures to ensure containment, such as isolating the patient and washing sheets with bleach.
Once rare, the viruses are now reemerging with increasing frequency, and have caused at least four outbreaks among humans in the last two years. Although the viruses are found most often in Africa, they have been unintentionally imported into the United States and Europe several times, and in recent years a version of the Ebola virus has been found replicating in swine raised for human consumption in Asia.
To conduct the study, Saphire and her group at TSRI collaborated with Yoshihiro Kawaoka, who holds joint appointments at the University of Wisconsin and University of Tokyo. Kawaoka’s group provided cellular microscopy and critical replication experiments to complement the TSRI team’s expertise in x-ray crystallography and protein biochemistry.
The results, five years in the making, revealed the Ebola VP40 protein exists as a dimer, not as a monomer as previously thought, and it rearranges its structure to assemble filaments to build the virus shell or “matrix” to release countless new viruses from infected cells. The study showed the protein also rearranges itself into rings in order to bind RNA and control the internal components of the virus copied inside infected cells.
This “shape-shifting” or “transformer” behavior explains how the Ebola virus can control a multi-step viral lifecycle using only a very limited number of genes.
In addition to Saphire, Bornholdt and Kawaoka, authors of the study, titled, “Structural basis for ebolavirus matrix and budding; protein plasticity allows multiple functions,” included Takeshi Noda, Dafna M. Abelson, Peter Halfmann and Malcolm R. Wood.
The research was supported by the Burroughs Welcome Fund, The Skaggs Institute of Chemical Biology at TSRI, the National Institutes of Health’s National Institute of Allergy and Infectious Diseases (numbers R43 AI1088843, 2T32AI007244 and U54 AI057153), the Japan Society for the Promotion of Science and the Japanese Ministry of Education, Culture, Sports, Science and Technology.
Send comments to: press[at]scripps.edu