The Russell Laboratory
Research
Cellular Responses to Double-Strand Breaks
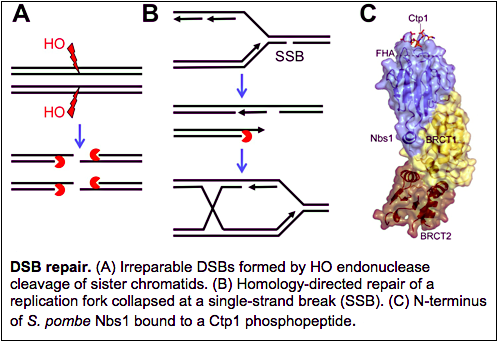 DNA double-strand breaks (DSBs) drive genome instability in cancer cells. They are a primary source of chromosome breaks that lead to chromosome translocations or loss of chromosome fragments. DSBs are an unavoidable consequence of normal cellular growth and metabolism, as they can arise spontaneously by collapse of replication forks, or by genetically programmed events in meiosis or V(D)J recombination during immune system development. DSBs can also arise from exogenous sources, notably cancer chemotherapeutics such as derivatives of etoposide or camptothecin, which are topoisomerase poisons that collapse replication forks, or by targeted ionizing radiation (IR), a commonly used anti-tumor therapy.
DNA double-strand breaks (DSBs) drive genome instability in cancer cells. They are a primary source of chromosome breaks that lead to chromosome translocations or loss of chromosome fragments. DSBs are an unavoidable consequence of normal cellular growth and metabolism, as they can arise spontaneously by collapse of replication forks, or by genetically programmed events in meiosis or V(D)J recombination during immune system development. DSBs can also arise from exogenous sources, notably cancer chemotherapeutics such as derivatives of etoposide or camptothecin, which are topoisomerase poisons that collapse replication forks, or by targeted ionizing radiation (IR), a commonly used anti-tumor therapy.
We are studying the proteins that detect, signal and repair DSBs. Key amongst these is a large molecular machine known as the Mre11-Rad50-Nbs1 (MRN) protein complex, often referred to as a “first-responder” to DSBs. Acting with another DSB repair protein known as CtIP or Ctp1, this complex “cleans up” DNA ends that may be covalently or non-covalently attached to proteins such as topoisomerases or Ku complex, and then initiates 5’-3’ resection that generates single-strand DNA overhangs required for homology-directed repair. The MRN complex simultaneously recruits ATM/Tel1 protein kinase, which initiates a signaling cascade that regulates DSB repair and arrests cell cycle progression. Mutations in the MRN subunits or its associated factors cause genome instability syndromes typified by neurological and immunological deficiencies, and often cancer. The MRN complex and ATM/Tel1 also have important roles at telomeres.
Protection of Genome Stability During DNA Replication
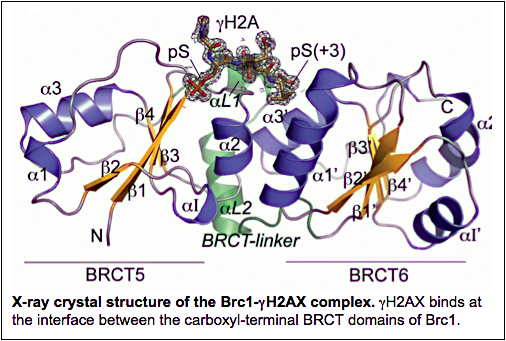 Genomes are especially vulnerable during the DNA synthesis (S) phase of the cell cycle, as replications forks frequently stall or collapse, often leading to the formation of one-ended DSBs. Damaged replication forks are recognized by ATR kinase, which phosphorylates a number critical substrates, including the carboxyl tail of histone H2AX in nucleosomes flanking the damaged fork. Phospho-histone H2AX, also known as γH2AX, converts chromatin into a recruitment platform for critical DNA damage response proteins. These include Brc1, which has six BRCT domains that were first identified in the BRCA1 breast cancer gene. By coating chromatin near the broken replication fork, Brc1 facilitates the reestablishment of an intact fork by homology-directed repair. This fork recapture mechanism generates and 4-way DNA junction known as a Holliday junction that is subsequently resolved by Mus81-Eme1 resolvase.
Genomes are especially vulnerable during the DNA synthesis (S) phase of the cell cycle, as replications forks frequently stall or collapse, often leading to the formation of one-ended DSBs. Damaged replication forks are recognized by ATR kinase, which phosphorylates a number critical substrates, including the carboxyl tail of histone H2AX in nucleosomes flanking the damaged fork. Phospho-histone H2AX, also known as γH2AX, converts chromatin into a recruitment platform for critical DNA damage response proteins. These include Brc1, which has six BRCT domains that were first identified in the BRCA1 breast cancer gene. By coating chromatin near the broken replication fork, Brc1 facilitates the reestablishment of an intact fork by homology-directed repair. This fork recapture mechanism generates and 4-way DNA junction known as a Holliday junction that is subsequently resolved by Mus81-Eme1 resolvase.
Cellular Responses to Environmental Toxicants
Heavy metals and metalloids such as cadmium, arsenic and hexavalent chromium are amongst the most widely distributed and dangerous environmental toxicants.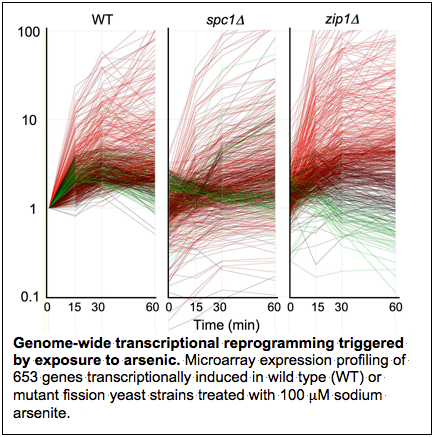 These toxins are linked to a wide range of diseases and they are of special concern for cancer, kidney damage, dementia, and birth defects. Detoxification mechanisms involving metal chelation have been defined in model organisms and human cells, but we lack a comprehensive picture about how cells mobilize and coordinate multiple detoxification, stress response and genome protection mechanisms in response to heavy metal exposure. We are using genetics, biochemistry, functional profiling of mutant libraries with next generation sequencing platforms, and mass spectrometry-based proteomics to investigate how the fission yeast Schizosaccharomyces pombe copes with exposure to heavy metals.
These toxins are linked to a wide range of diseases and they are of special concern for cancer, kidney damage, dementia, and birth defects. Detoxification mechanisms involving metal chelation have been defined in model organisms and human cells, but we lack a comprehensive picture about how cells mobilize and coordinate multiple detoxification, stress response and genome protection mechanisms in response to heavy metal exposure. We are using genetics, biochemistry, functional profiling of mutant libraries with next generation sequencing platforms, and mass spectrometry-based proteomics to investigate how the fission yeast Schizosaccharomyces pombe copes with exposure to heavy metals.