Social distancing was more effective at preventing local COVID-19 transmission than international border closures
Scripps Research scientists and collaborators uncovered the relative effectiveness of different COVID-19 mandates, helping guide public health policy for future viral threats.
December 14, 2023
LA JOLLA, CA—Elucidating human contact networks could help predict and prevent the transmission of SARS-CoV-2 and future pandemic threats. A new study from Scripps Research scientists and collaborators points to which public health protocols worked to mitigate the spread of COVID-19—and which ones didn’t.
In the study, published online in Cell on December 14, 2023, the Scripps Research-led team of scientists investigated the efficacy of different mandates—including stay-at-home measures, social distancing and travel restrictions—at preventing local and regional transmission during different phases of the COVID-19 pandemic. They found that local transmission was driven by the amount of travel between locations, not by how geographically nearby they were. The study also revealed that the partial closure of the U.S.-Mexico border was ineffective at preventing cross-border transmission of the virus. These findings, in combination with ongoing genomic surveillance, could help guide public health policy to prevent future pandemics and mitigate the new “endemic” phase of COVID-19.
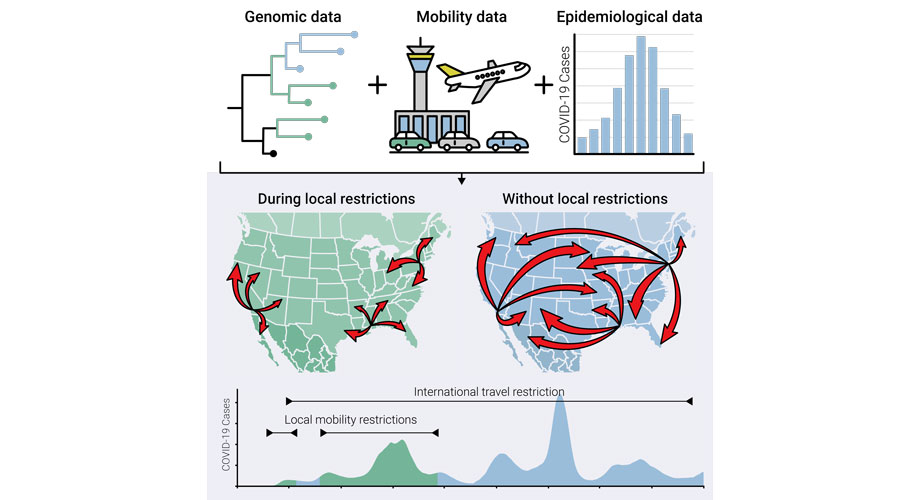
“We show that it’s not necessarily how geographically close locations are to each other; the real measure is how connected two locations are in terms of the movement of people,” says co-senior author Mark Zeller, PhD, a project scientist and genomic epidemiologist in Kristian Andersen’s lab at Scripps Research. “SARS-CoV-2 is probably a good model for respiratory virus spread in general, and knowing how these viruses are spreading can help us develop targeted measures for future pandemics.”
The study was led by Scripps Research scientists in collaboration with the County of San Diego, UCSD and various academic laboratories, public health laboratories and hospitals in California and Baja California, Mexico.
“This invaluable information helps direct strategies to mitigate the spread of future viral threats—including our ongoing genomic surveillance methods, such as wastewater testing,” says Seema Shah, MD, medical director of San Diego County’s epidemiology and immunization services branch. “In an era where global challenges require united responses, this study is also a testament to the close-knit collaboration between academic research institutions and public health laboratories and hospitals on both sides of the border.”
To explain how changing travel patterns impacted COVID-19 transmission during the first two years of the pandemic, the team quantified the genetic similarity between viral sequences collected at different locations and points in time.
“The more similar the virus genomes are between two different locations, the more connected those locations are in terms of the movement of people,” says Zeller.
The researchers sequenced more than 82,000 SARS-CoV-2 samples from San Diego, California and Baja California, Mexico and compared these sequences to SARS-CoV-2 genomes from North America and the rest of the world. Then they integrated the genomic data with a database of air and land travel data (based on anonymous cellphone tracking) to estimate the connectivity of different counties within North America.
During the early phases of the pandemic, when COVID-19 mandates included strict stay-at-home and social distancing measures, transmission mainly occurred at the local and regional level—within and between neighboring counties. But as these mandates were relaxed and people began to travel more freely, predictably, transmission from distant locations increased.
“Mobility between neighboring counties remained almost constant throughout the entire first two years of the pandemic, but it was these long-distance movements, from San Diego to more distant counties or to neighboring states, that were really disrupted by the early mandates,” says genomic epidemiologist and first author Nathaniel Matteson, PhD, who worked on the study as a graduate student in the Andersen Lab at Scripps Research. “After the mandates were relaxed, about 50% of the virus circulating in San Diego was the result of locally circulating virus, and 50% had been recently introduced as the result of travel between locations.”
The analysis also revealed there was continuous transmission between San Diego and nearby Baja California during the entire duration of the pandemic, suggesting that the partial closure of the U.S.-Mexico border was ineffective in preventing transmission.
“This finding adds to this growing body of evidence that these targeted measures that try to stop the flow of people from one specific location are quite ineffective, whereas social distancing measures have shown to be much more effective at reducing transmission,” says Matteson. “It also highlights the importance of regional and international collaborations and resource sharing for effective disease surveillance and prevention.”
Next, the researchers plan to study transmission dynamics at an even finer scale. “Now we’re looking at hyper-local transmission within San Diego County—for example how transmission is influenced by things like commuting patterns,” says Zeller.
In addition to Zeller and Matteson, authors of the study, “Genomic surveillance reveals dynamic shifts in the connectivity of COVID-19 epidemics,” include Ezra Kurzban, Madison A. Schwab, Sarah A. Perkins, Karthik Gangavarapu, Joshua I. Levy, Edyth Parker, Manuel Sanchez-Alavez, Alana Weiss, Catelyn Anderson, Christine M. Aceves, Emily G. Spencer, Alison J. King, Emory C. Hufbauer, Justin J. Lee, Karthik S. Ramesh, Kelly N. Nguyen, Kieran Saucedo, Refugio Robles-Sikisaka, Laura Nicholson, Kristian G. Andersen, and Shirlee Wohl of Scripps Research.
This work was supported by funding from the National Institutes of Health (U19AI135995, U01AI151812, UL1TR002550, F32AI154824, and R01AI153044), The Prebys Foundation, and the CDC contract 7530122C14843. Computing resources for this research were donated by NVIDIA Corporation and Advanced Micro Devices, Inc.
For more information, contact press@scripps.edu

