Research
Structure, Dynamics and Interactions of Proteins
My laboratory utilizes high-resolution nuclear magnetic resonance (NMR) spectroscopy and other biophysical and biochemical methods to investigate the structure, dynamics, and folding mechanisms of proteins and to map their functional interactions. NMR is unique as a method for determining three-dimensional structures of proteins and protein complexes in solution and also providing novel information about the time-dependent structural fluctuations that are essential for protein function.
Intrinsically disordered proteins and cellular signaling.
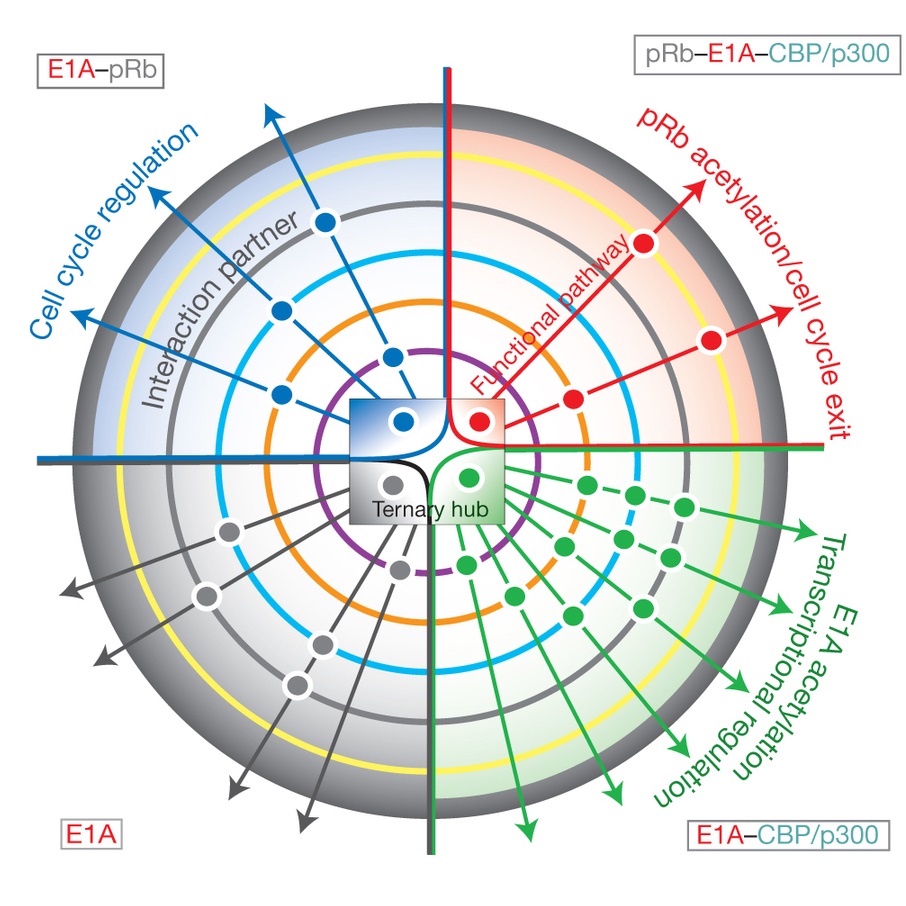
Intrinsically disordered proteins are highly abundant in eukaryotes and play a central role in cellular regulatory processes and signaling pathways. We are using a multidisciplinary approach, including a broad range of biochemical and biophysical methods, NMR, and single molecule fluorescence (in collaboration with Ashok Deniz), to elucidate the structure of the general transcriptional coactivators CBP and p300 and characterize their functional interactions with key cellular and viral targets. We are implementing novel NMR methods, intein labeling technologies, and single molecule FRET methods to characterize the structure of disordered proteins and their complexes and to elucidate the mechanism by which disordered proteins fold upon binding to their targets.
Intrinsically disordered proteins are highly abundant in eukaryotes and play a central role in cellular regulatory processes and signaling pathways. We are using a multidisciplinary approach, including a broad range of biochemical and biophysical methods, NMR, and single molecule fluorescence (in collaboration with Ashok Deniz), to elucidate the structure of the general transcriptional coactivators CBP and p300 and characterize their functional interactions with key cellular and viral targets. We are implementing novel NMR methods, intein labeling technologies, and single molecule FRET methods to characterize the structure of disordered proteins and their complexes and to elucidate the mechanism by which disordered proteins fold upon binding to their targets.
Mechanisms of nucleic acid recognition by zinc finger proteins.
We are using NMR and X-ray crystallography (in collaboration with Ian Wilson’s laboratory) to elucidate the structural basis by which the protein Muscleblind recognizes pathogenic RNA sequences, and by which the protein Kaiso binds both regulatory and methylated DNA motifs.
We are using NMR and X-ray crystallography (in collaboration with Ian Wilson’s laboratory) to elucidate the structural basis by which the protein Muscleblind recognizes pathogenic RNA sequences, and by which the protein Kaiso binds both regulatory and methylated DNA motifs.
Mechanisms of protein folding and misfolding.
NMR is uniquely suited for studies of protein folding and misfolding pathways, providing detailed insights into the structure and dynamics of unfolded states and partially folded intermediates. We are applying NMR relaxation dispersion methods to elucidate the molecular mechanism by which the protein transthyretin spontaneously unfolds and aggregates, leading to amyloid disease.
NMR is uniquely suited for studies of protein folding and misfolding pathways, providing detailed insights into the structure and dynamics of unfolded states and partially folded intermediates. We are applying NMR relaxation dispersion methods to elucidate the molecular mechanism by which the protein transthyretin spontaneously unfolds and aggregates, leading to amyloid disease.
Protein dynamics and “invisible” excited states.
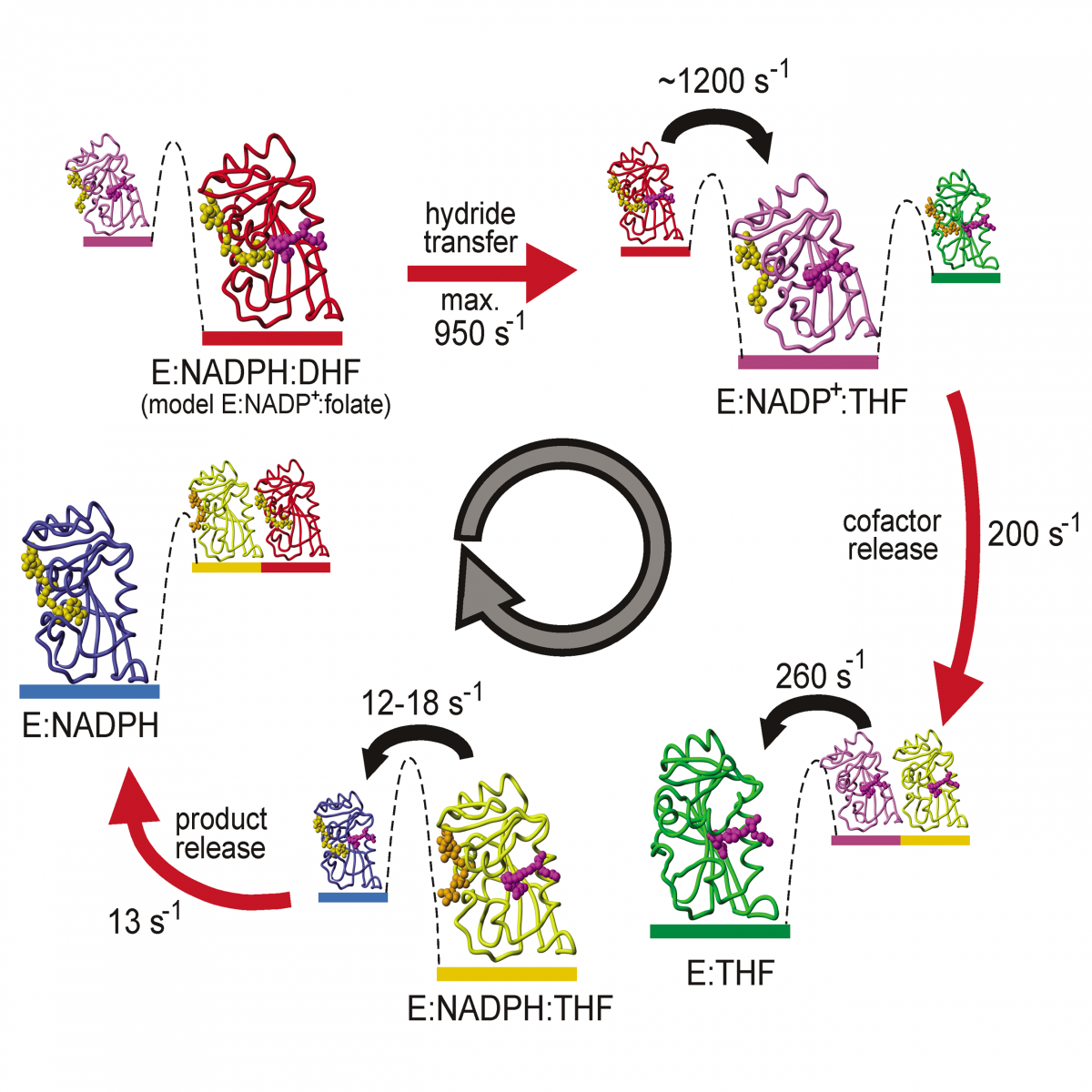
We are applying NMR relaxation dispersion methods to characterize the dynamics of the enzyme dihydrofolate reductase and to determine the structure of weakly populated excited states that play a functional role in catalysis. These studies, which are also being applied to study the dynamics of a stress-activated protein kinase, are providing unprecedented insights into the intrinsic dynamics of enzymes and their role in catalysis.
We are applying NMR relaxation dispersion methods to characterize the dynamics of the enzyme dihydrofolate reductase and to determine the structure of weakly populated excited states that play a functional role in catalysis. These studies, which are also being applied to study the dynamics of a stress-activated protein kinase, are providing unprecedented insights into the intrinsic dynamics of enzymes and their role in catalysis.
Research Techniques
NMR Spectroscopy
X-ray Crystallography
Mass Spectrometry
Fluorescence based methods
Isothermal Titration Calorimetry
Computational tools
Image Gallery
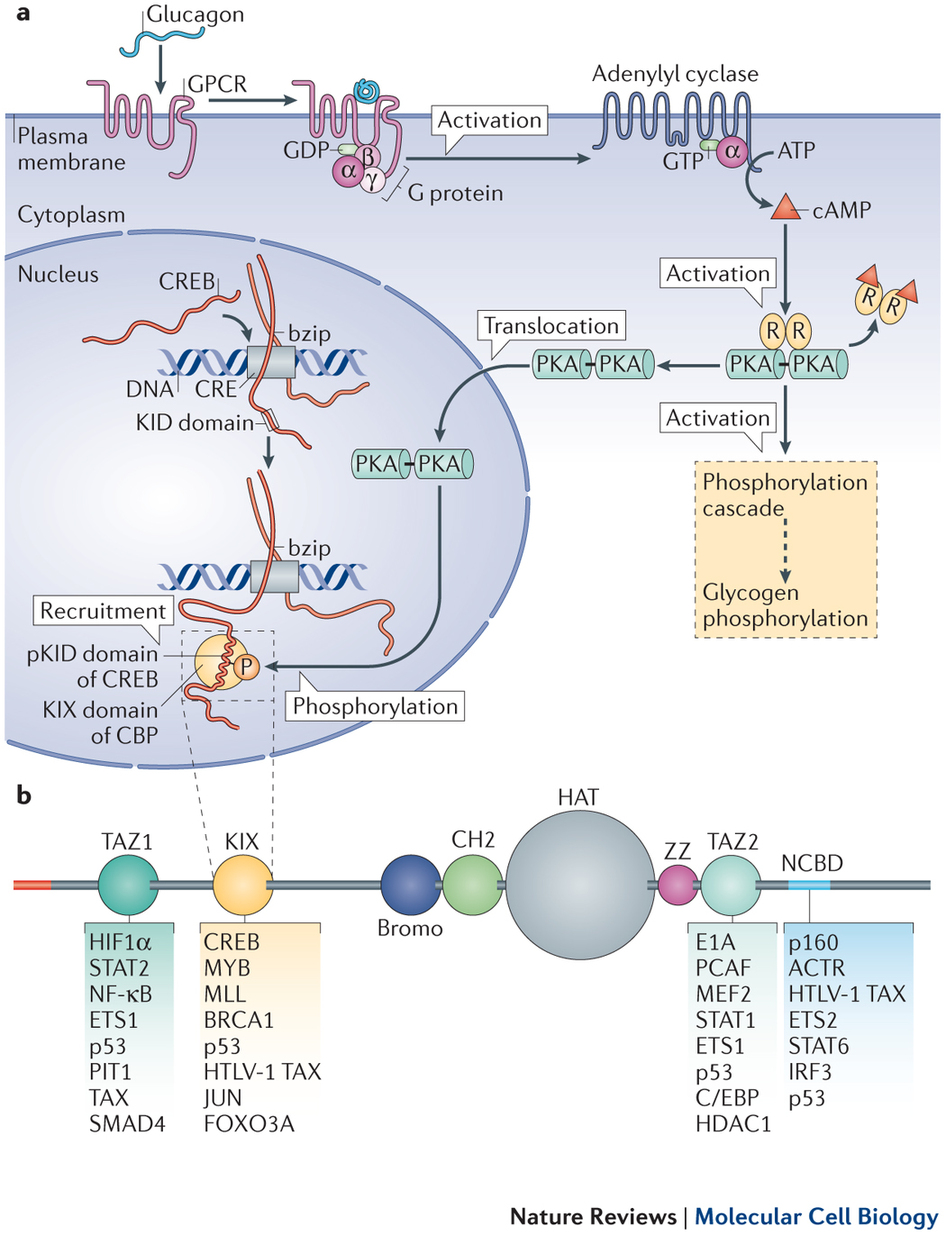
Read more: Nature Reviews Molecular Cell Biology
Intrinsically disordered proteins in cellular signalling and regulation.
Read more: PNAS
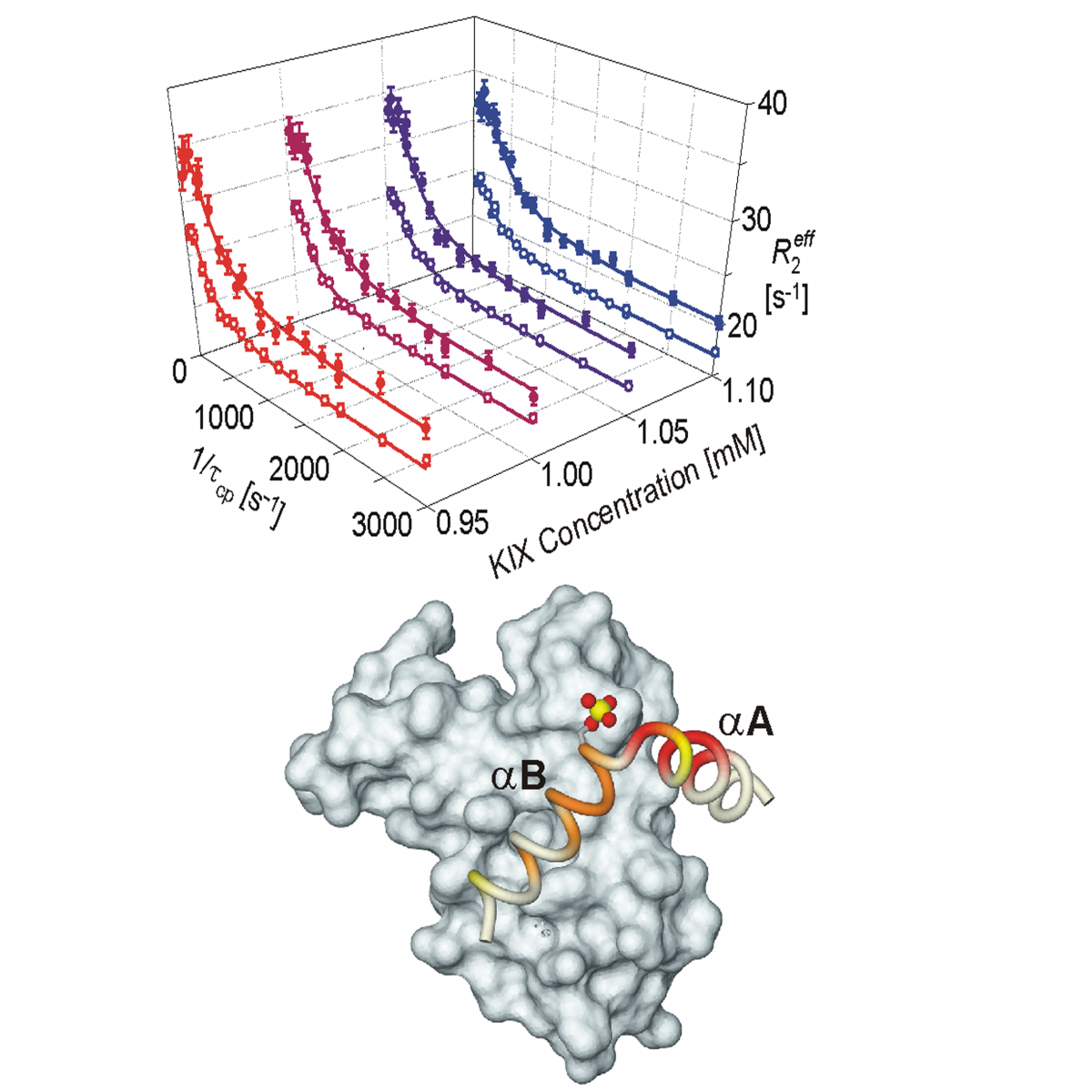
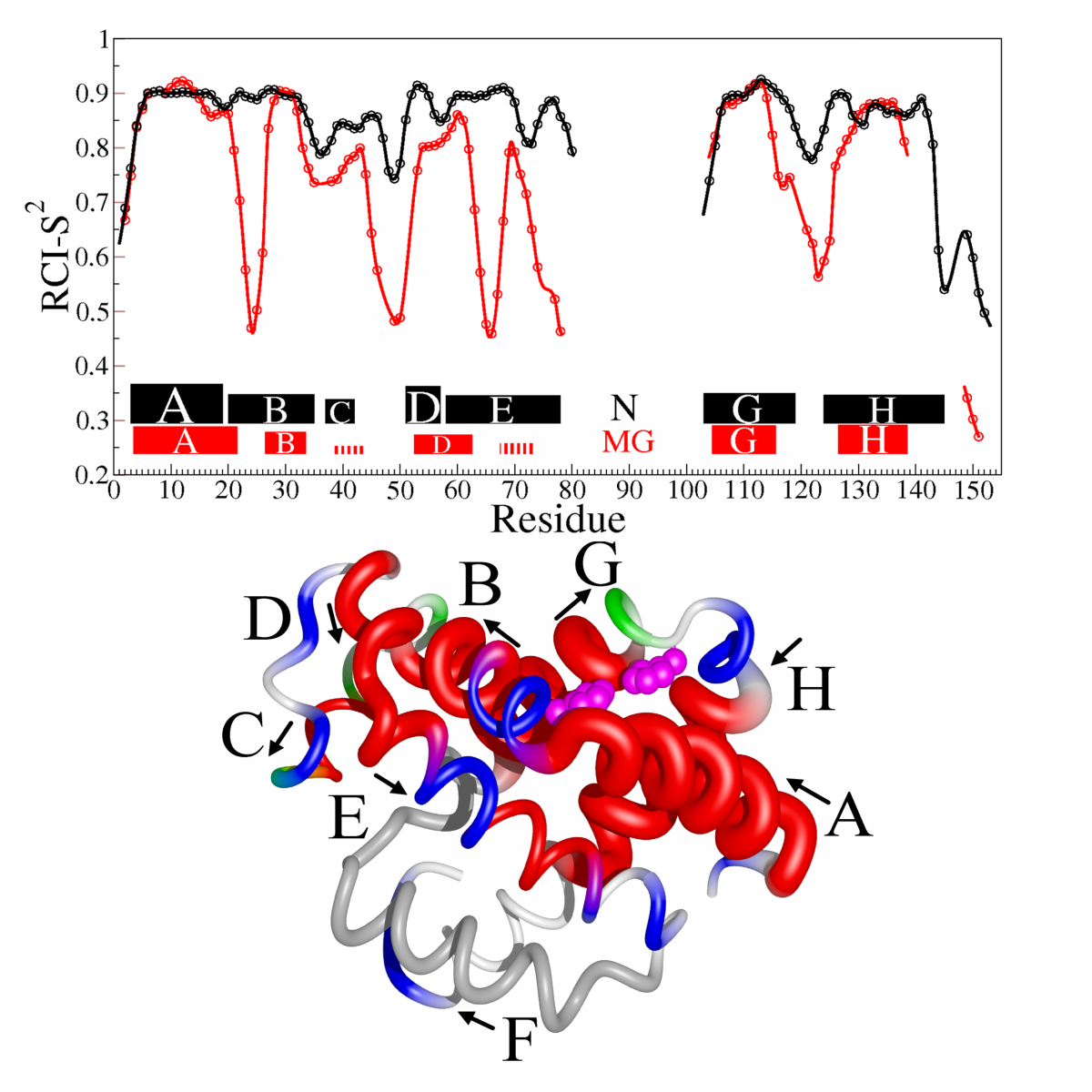
Measurement of protein unfolding/refolding kinetics and structural characterization of hidden intermediates by NMR relaxation dispersion.
Read more: PNAS
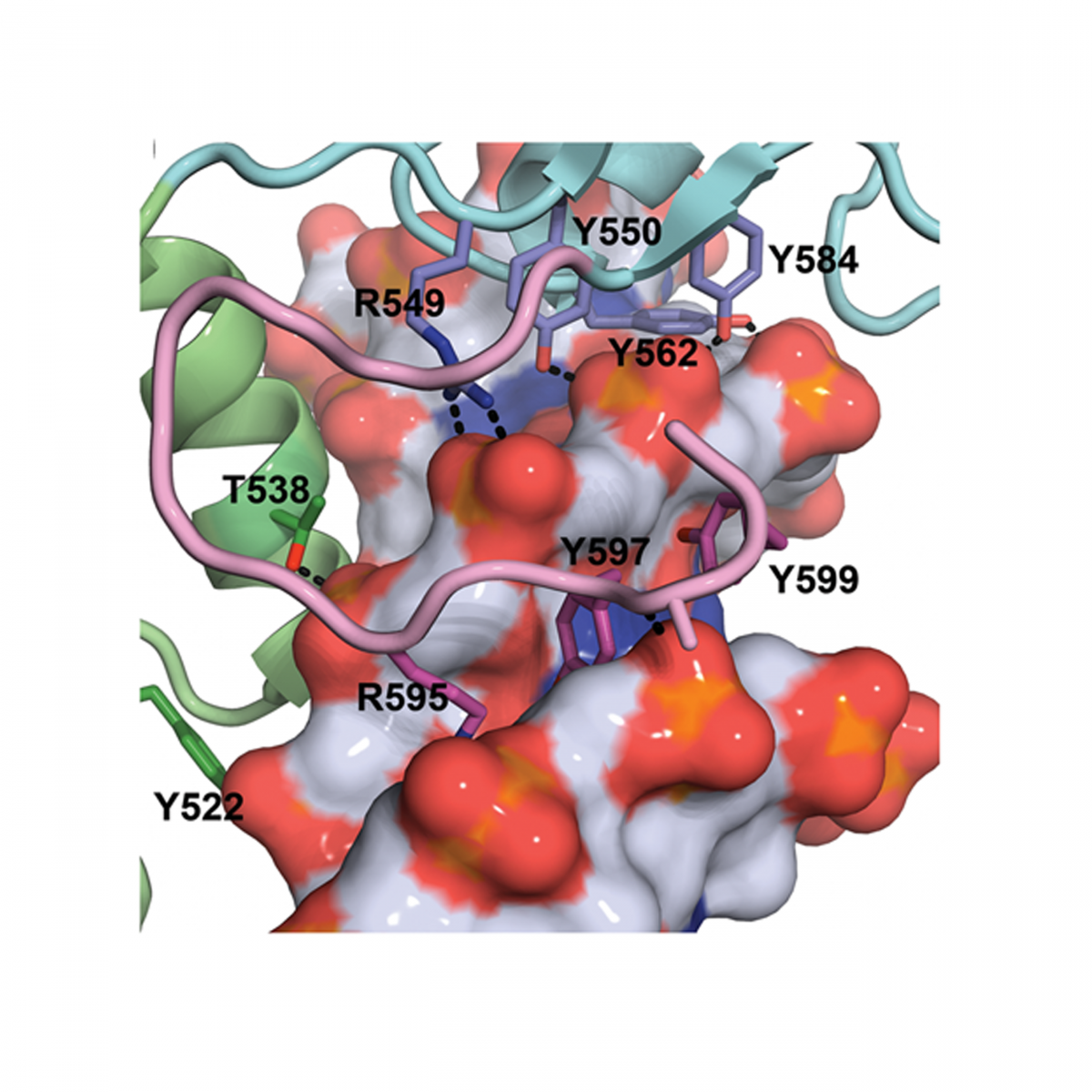
Molecular basis for recognition of methylated and specific DNA sequences by the zinc finger protein Kaiso.