Crucial Genetic Diversity Enzyme Long Sought by Biologists
Discovered by Scientists at The Scripps Research Institute
By Jason Socrates
Bardi
Simultaneous reports by two teams, led by Professor Paul
Russell and Associate Professor Clare H. McGowan, both of
The Scripps Research Institute (TSRI), identify the "resolvase"
enzyme that may be responsible for generating genetic diversity
during sexual reproduction. The discovery of the enzyme could
lead to improved cancer chemotherapy.
In the current issues of the journals Cell and Molecular
Cell, the researchers have published papers that describe
Mus81, a resolvase enzyme of the fission yeast Schizosaccharomyces
pombe, and its human analog.
Resolvase is essential for a crucial step in DNA recombination,
says Russell, because it is the molecule that allows two chromosomes
to cross over. "It is one of the most important enzymes involved
in genetic recombination," he says.
Genetic recombination occurs in the process of meiosis,
when chromosomes from the mother and father become paired.
The aligned chromosomes break and DNA strands from both chromosomes
become intertwined at the point of the cross-over. At the
molecular level, this combining happens at what is called
a "Holliday junction," where the strands of DNA literally
cross one another.
However, the DNA must at some point be uncrossed by cutting
across the Holliday junction in the last crucial step in genetic
recombination. This is the responsibility of resolvase enzymes.
The final product of this process is a pair of new chromosomes
that have genetic material from both parents.
"[Resolvase] is the molecule that allows children to inherit
a unique mixture of traits from mother and father, without
it we wouldn't have the infinite range of genetic combinations
that makes us all different," says McGowan.
It has long been known that there should be such enzymes,
and several examples from other organisms, such as bacteria,
have been around for years. And for years, scientists have
searched for the resolvase gene in eukaryotic cells, such
as humans and yeast, which have linear chromosomes packaged
in a nucleus. Until now, none has been found.
Russell and his colleagues showed that Mus81 is an essential
component of the resolvase enzyme in yeast cells. Mus81 is
structurally unrelated to bacterial resolvases. In a related
work, McGowan's study demonstrated that a human analog of
the Mus81 protein also has resolvase activity.
The identification of a human resolvase may have a profound
effect on cancer therapy because the enzyme also has an important
role in cell replication.
When cells are replicating their DNA prior to division,
they have mechanisms to sense if the DNA is damaged. When
the DNA is damaged, a cell's replication machinery will stop,
spontaneously back up and form a Holliday junction. Resolvase
recombines DNA strands at Holliday junctions and this allows
the replication machinery to bypass the damaged DNA.
Cancer cells are often defective in the mechanisms that
sense damaged DNA. Russell and McGowan envision that treatment
of tumors with chemotherapeutics that damage DNA, combined
with rational targeting of resolvase activity, could be a
highly potent cancer treatment.
This research is also another vindication of fission yeast
as a model organism for human biology. Resembling humans,
S. pombe cells reproduce sexually through meiosis and
have a similar cell cycle. Because of the ease of manipulating
yeast genetically and because of their uncanny similarities,
S. pombe is a good model system for studying the human
cell cycle.
"S. pombe has contributed enormously towards understanding
the human cell cycle and towards advances in the treatment
and understanding of cancer," says McGowan.
The research article "Mus81-Eme1 Are Essential Components
of a Holliday Junction Resolvase" is authored by Michael N.
Boddy, Pierre-Henri L. Gaillard, W. Hayes McDonald, Paul Shanahan,
John R. Yates 3rd, and Paul Russell and appears in the November
16, 2001 issue of Cell.
The research article "Human Mus81-Associated Endonuclease
Cleaves Holliday Junctions In Vitro" is authored by Xiao-Bo
Chen, Roberta Melchionna, Cecile-Marie Denis, Pierre-Henri
L. Gaillard, Alessandra Blasina, Inez Van de Weyer, Michael
N. Boddy, Paul Russell, Jorge Vialard, and Clare H. McGowan
and appears in the November, 2001 issue of Molecular Cell.
The research was funded by the National Institutes of Health,
by The R.W. Johnson Pharmaceutical Research Institute, and
by the Janssen Research Foundation.

|

Professor Paul Russell (left) recently
published his work on the enzyme resolvase in the journal
"Cell." Associate Professor Clare H. McGowan (right)
recently published a separate paper on the enzyme in the journal
"Molecular Cell."
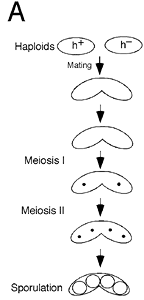
Diagrammatic representation of meiosis
in fission yeast. Cells of opposite mating type (h+/h–)
mate and undergo replication followed by two nuclear divisions,
resulting in haploid spores.
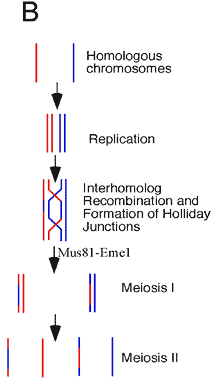
During meiosis homologous chromosomes
are replicated, followed by recombination between homologous
chromosomes. Double Holliday junctions are formed as a result
of recombination and must be cleaved to allow segregation
of chromosomes during the first nuclear division, or meiosis
I. In fission yeast this important task is performed by a
resolvase called Mus81-Eme1.
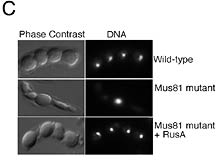
Microscopic examination of fission yeast
meiotic products. The top panels show the four haploid spores
(phase contrast) and nuclei (DNA) expected from meiosis in
fission yeast. The middle panels show the defect in meiosis
observed in Mus81 mutants. Unresolved Holliday junctions prevent
segregation of chromosomes. This defect results in the formation
of one large spore containing all the DNA and some smaller
spores with no nuclei. The lower panels show that RusA, a
bacterial Holliday junction resolvase, is able to substitute
for the function of Mus81-Eme1 in meiosis.
|

