- Romesberg Lab
- Research
- Publications
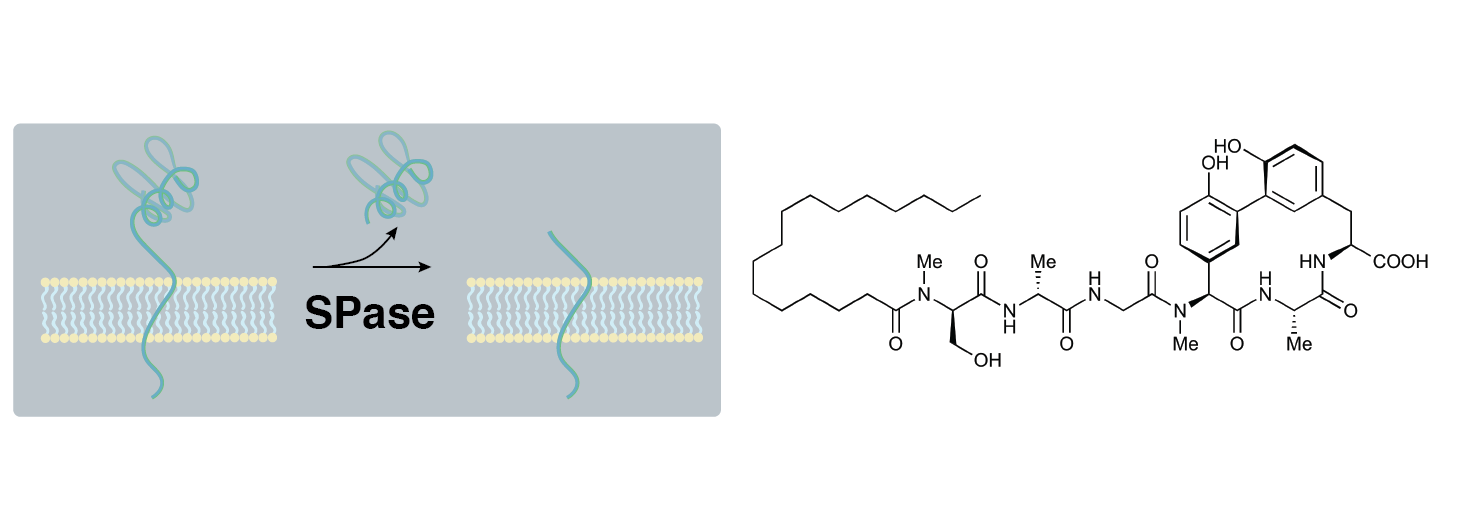
The arylomycins inhibit SPase, which is required for protein secretion - but many bacteria have naturally evolved resistance. We're developing arylomycins that overcome this resistance and looking for other latent antibiotics.
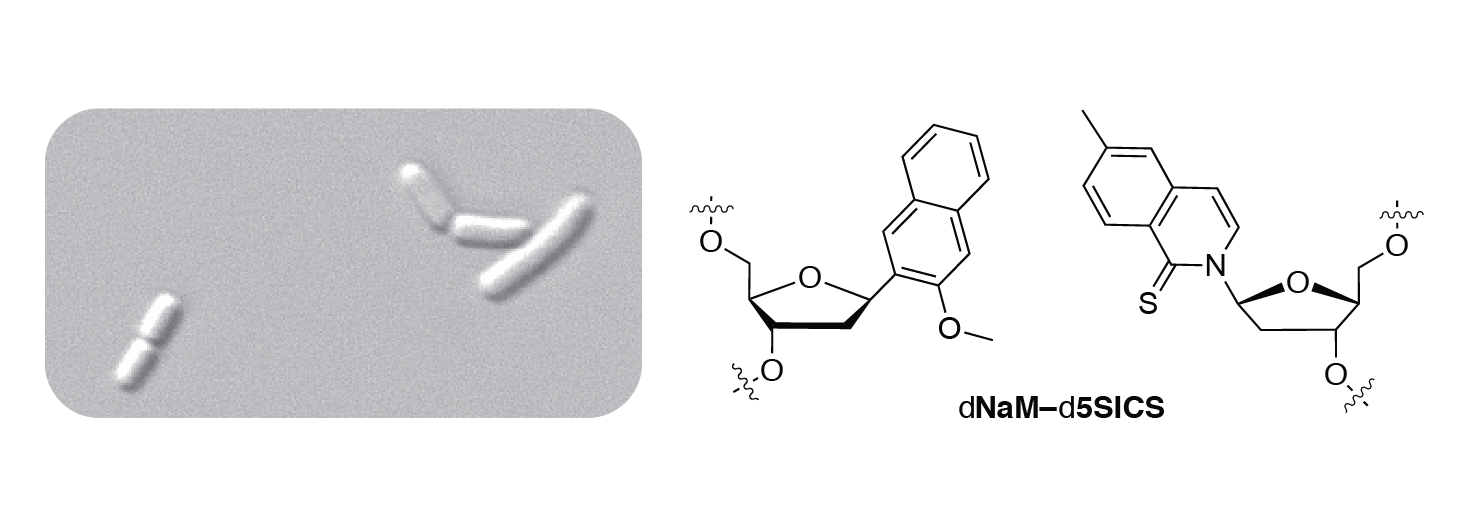
We've surpassed our longstanding goal of developing a replicable unnatural DNA base pair and are now focused on not just storage, but also retrieval of information in an expanded genetic alphabet in vivo.
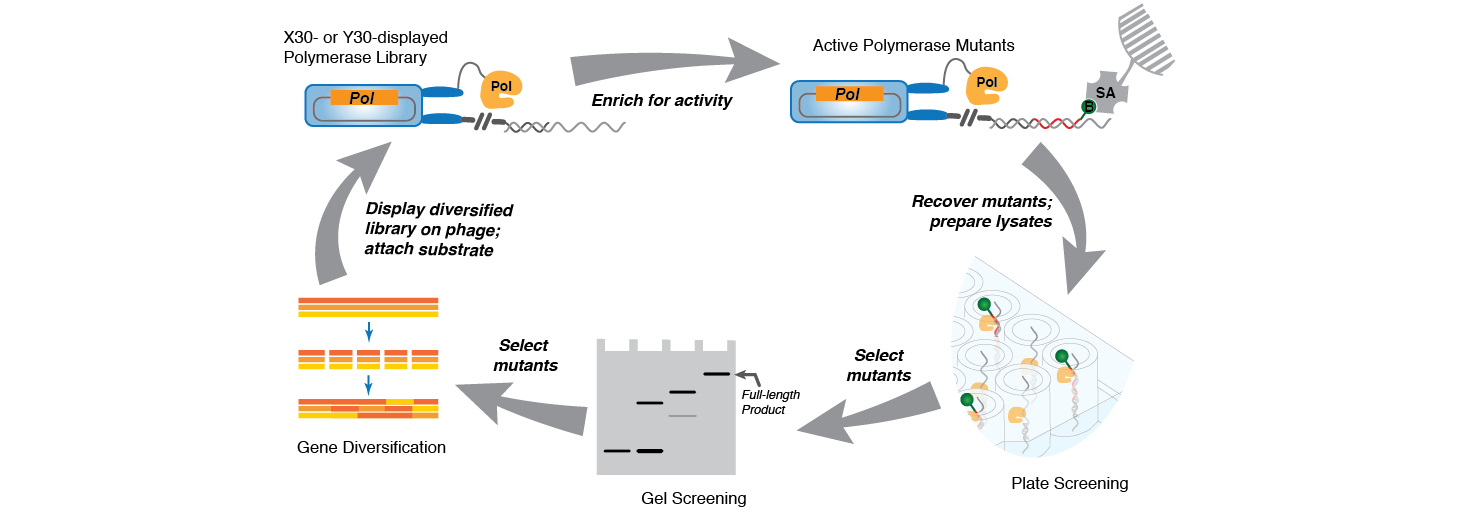
We've evolved polymerases that accept non-natural substrates, e.g. unnatural or modified natural R/DNA. Recent work has focused on applying this progress to the evolution of nuclease-resistant aptamers.
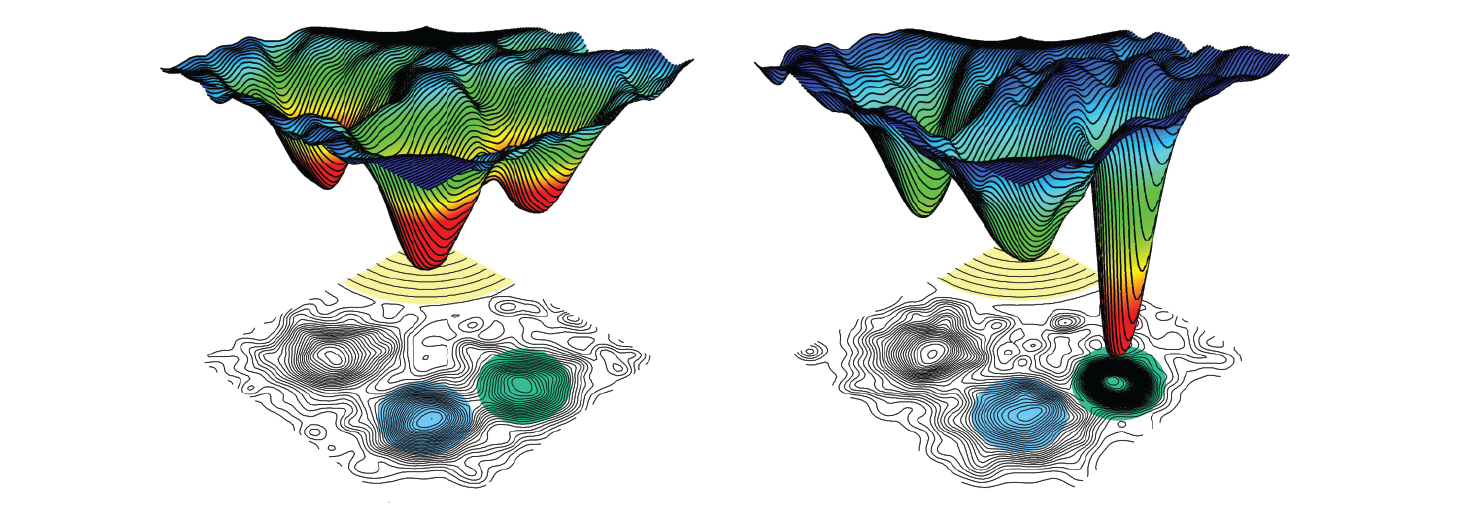
We're characterizing antibodies at different stages of their evolution to examine how somatic mutations affect dynamics and function. We're also developing the use of carbon-deuterium bonds as general probes of protein dynamics.
reading suggestions for busy people
Zimmermann, PNAS 2006
Zhang, PNAS 2017
Adhikary, JACS 2014
Craney, mBio 2015