Rethinking Tau
By Jason Socrates Bardi
Scientists would do well not to listen to the wisdom
of famous jazz legend Louis Armstrong, who when asked to define
jazz music, supposedly replied, if you have to ask,
you'll never know. For scientists, better advice would be
if you don't ask you'll never know.
One of the unanswered questions in the field of neuroscience
has been how neurons in the brain develop and form connections—sometimes
as many as 10,000 apiece. This question is of great interest
to scientists because neurons are irreversibly damaged or
lost in spinal cord injuries and neurodegenerative diseases
like Alzheimer's.
Associate Professor Shelley Halpain, Research Associate
Benoit Roger, and several of their colleagues at The Scripps
Research Institute report progress in this area in a recent
article in the journal Current Biology.
Similar Structures, Different Results
What the researchers were asking in particular was how two
different neuronal proteins help maturing neurons send out
neurites—the long finger-like processes characteristic
of mature neurons that connect them with other neurons.
As these neurites are forming, they must be supported by
the cell's cytoskeleton—its actin filaments and microtubules—which
means that for the proper formation of the neurites, the microtubules
and actin filaments must assemble at the same time. In recent
years, scientists have also begun to appreciate that microtubules
and actin filaments must interact with each other during this
process. Scientists have identified a number of proteins that
mediate this interaction, including the microtubule-associated
proteins MAP2 and tau.
MAP2 and tau are abundant in neurons where they stabilize
and promote the growth of the microtubules—something
needed for neurite outgrowth.
Roger and Halpain's experiments showed that MAP2 also binds
to actin. The results showed that the domain of MAP2 that
binds to actin is the same domain that binds to the microtubules.
In contrast, the similar domain on the tau protein, which
also binds microtubules, does not bind to actin the same way.
In fact, tau has no actin binding at all.
This was a surprise because MAP2 and tau are so similar
structurally—67 percent of the amino acid of the implicated
cytoskeleton binding domain sequences are identical, and they
both bind to microtubules with almost the same activity. However,
Roger and Halpain found that MAP2 is sufficient to trigger
neuritic growth but tau is not. And by making a "chimeric"
protein of tau with one piece of MAP2 exchanged (the piece
that binds to actin), Roger and Halpain showed that this altered
tau could now induce neurites.
These differences between MAP2 and tau may cause scientists
to rethink the role of tau in neurons and in various neurological
disorders. For a long time, scientists have known that tau
protein form abnormal aggregates inside cells in Alzheimer's
disease, even though the amyloid proteins that form plaques
outside of cells were thought to be the actual cause of the
disease. Nevertheless, several other diseases are now known
to result directly from defective tau—these are called
the tauopathies. These rare hereditary dementias, which were
just discovered in the last decade, are caused by single amino
acid mutations in tau that cause the protein to form fibrous
"neurofibrillary" tangles inside neurons.
Interestingly, no such mutations have been found to cause
the MAP2 protein to form tangles. Perhaps the ability of MAP2
to interact with actin as well as microtubules may prevent
it from forming neurofibrillary tangles. Such information
may be used in the future to determine how altering tau's
structure could prevent neurodegenerative diseases.
To read the article, "MAP2c, but Not Tau, Binds and Bundles
F-Actin via Its Microtubule Binding Domain" by Benoit Roger,
Jawdat Al-Bassam, Leif Dehmelt, Ronald A. Milligan, and Shelley
Halpain, see the March 9, 2004 issue of Current
Biology.
Send comments to: jasonb@scripps.edu

|
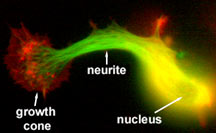
Neuronal cell in the process of initiating
a neurite. Microtubules are shown in green, actin filaments
in red. Image courtesy of Leif Dehmelt, Halpain lab.
|

