

Tweet
Scientists Design New Anti-Flu Virus Proteins Using Computational Methods
A research article in the May 12, 2011 issue of the journal Science demonstrates the use of computational methods to design new antiviral proteins not found in nature, but capable of targeting specific surfaces of flu virus molecules.
One goal of such protein design would be to block molecular mechanisms involved in cell invasion and virus reproduction.
Computationally designed, antiviral proteins binding a particular surface on a target protein might also have diagnostic and therapeutic potential in identifying and fighting viral infections.
The first authors of the study are Damian C. Ekiert of The Scripps Research Institute and Sarel J. Fleishman and Timothy Whitehead of the University of Washington (UW). The senior authors are Ian Wilson, Hansen Professor of Structural Biology and member of the Skaggs Institute for Chemical Biology at Scripps Research, and David Baker, professor at the UW and Howard Hughes Medical Institute.
The researchers note that additional studies are required to see if such designed proteins can help in diagnosing, preventing or treating viral illness. What the study does suggest is the feasibility of using computer design to create new proteins with antiviral properties.
"Influenza presents a serious public health challenge," the researchers noted, "and new therapies are needed to combat viruses that are resistant to existing anti-viral medications or that escape the body's defense systems."
The scientists focused their attention on the section of the flu virus known as the hemagglutinin stem region. They concentrated on trying to disable this part because of its function in helping the virus invade the cells of the human respiratory tract.
The researchers' approach was somewhat similar to engineering a small space shuttle with the right configuration and construction, as well as recognizance and interlocking mechanisms, to dock with a troublesome space station and upset its mission. Only these scientists attempted their engineering feat at an atomic and molecular level.
Central to their approach is the ability of biological molecules to recognize certain other molecules or their working parts, and to have an affinity for binding to them at pre-determined locations. This recognition has both physical and chemical bases. Protein-protein interactions underlie many biological activities, including those that disarm and deactivate viruses.
In their report, the researchers described their general computational methods for designing new, tiny protein molecules that could bind to a certain spot on large protein molecules. They took apart some protein structures and watched how these disembodied sections interacted with a target surface. They analyzed particular high-affinity interactions, and used this information to further refine computer-generated designs for interfaces.
"Protein surfaces are never flat, but have many crevices and bulges at the atomic scale," Fleishman explained. "The challenge is to identify amino acid side chains that would fit perfectly into these surfaces. The fit must be precise both in shape and in other chemical properties such as electrostatic charge. This geometrical and biophysical problem can be computationally solved, but requires large computational resources."
The researchers made use of a peer-to-peer computing platform called Rosetta@Home for going through the hundreds of millions of possible interactions of designed proteins and the surface of the hemagglutinin to solve this challenge. Following optimization, the designed proteins bound the hemagglutinin very tightly. Through this method, the researchers created two designs for new proteins that could bind to a surface patch on the stem of the influenza hemagglutinin from the 1918 H1N1 pandemic flu virus. But because this surface is similar in many influenza viruses, the designed proteins can target a diverse set of viruses, including the 2009 H1N1 "swine flu" and the highly pathogenic H5N1 "bird flu."
The shortcomings of the approach, due to approximations, meant that the researchers started out with 73 possibilities of which just two were successful. One of the characteristics of the influenza hemagglutinin stem is that it changes shape by refolding when in an acidic environment to enable membrane fusion to occur and allow the viral RNA to get inside the cell and thereby allow the virus to reproduce itself inside human respiratory cells. In this study, one of the newly designed proteins was shown to block this conformational change, not only in H1 influenza hemagglutinin, but also in H5 avian influenza ("bird flu").
"This finding suggests that this new protein design may have virus-neutralizing effects against multiple influenza subtypes," the researchers reported.
What was unusual about the workable designs was that they had helical binding modes, roughly shaped like a spiral staircase, rather than the loop binding that naturally occurring antibodies employ.
X-ray crystallography of the protein complex showed that the actual orientation of the bound proteins was almost identical to the way the binding mode was designed. The modified surface of the main recognition helix on the designed protein was packed into a groove on the desired region of the virus protein.
"Overall, the crystal structure is in excellent agreement with the designed interface," the researchers noted, "with no significant deviations at any of the contact points." The design and the actual formation were nearly identical.
The scientists were encouraged by this finding as the design methods, the scientists believe, through iteration and selection, captured the essential features of the desired protein-protein interaction.
In addition to Fleishman, Whitehead, Damian C. Ekiert, Wilson, and Baker, authors of the study, "Computational Design of Proteins Targeting the Conserved Stem Region of Influenza Hemagglutinin," include Cyrille Dreyfus of Scripps Research and Jacob E. Corn and Eva-Maria Strauch of UW. For more information, see http://www.sciencemag.org/content/332/6031/816.abstract.
This research was supported by grants to the Baker lab from Defense Advanced Research Projects Agency, the National Institutes of Health (NIH) yeast resource center, the Defense Threat Reduction Agency, and the Howard Hughes Medical Institute, funding to the Wilson lab including a grant from NIH National Institute of Allergy and Infectious Diseases and predoctoral fellowships from the Achievement Rewards for College Scientists Foundation and the NIH Molecular Evolution Training Program, as well as funds from the Skaggs Institute for Chemical Biology. Additional funding was supplied by a fellowship from Human Frontier Science Program, the Jane Coffin Childs Memorial Fund, and a career development award from the NIH National Institute of Allergy and Infectious Diseases.
Computational designs were generated on resources generously provided by participants of Rosetta @ Home and the Argonne National Leadership Computing Facility. X-ray diffraction data sets were collected at the Stanford Synchrotron Radiation Lightsource beamline 9-2 and at the Advanced Photon Source (APS) beamline (GM/CA-CAT). The GM/CA CAT 23-ID-B was funded in whole or in part with federal funds from National Cancer Institute and the NIH National Institute of General Medical Science. Use of the APS was supported by the U.S. Department of Energy.
Send comments to: mikaono[at]scripps.edu
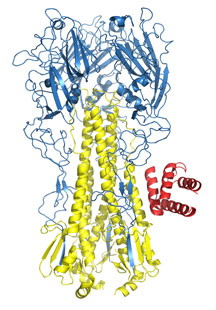 Members of the Wilson lab and colleagues from the University of Washington used computational methods to design new antiviral proteins capable of targeting specific surfaces of the flu virus. Here, a hemagglutinin spike (blue and yellow ribbons) from influenza A is shown bound to one of the new anti-viral proteins, HB36 (red). (Image courtesy of the Wilson lab. Click for details.)
Members of the Wilson lab and colleagues from the University of Washington used computational methods to design new antiviral proteins capable of targeting specific surfaces of the flu virus. Here, a hemagglutinin spike (blue and yellow ribbons) from influenza A is shown bound to one of the new anti-viral proteins, HB36 (red). (Image courtesy of the Wilson lab. Click for details.)
