

Tweet
Structure of DNA Repair Complex Reveals Workings of Powerful Cell Motor
By Renee Twombly
Over the last years, two teams of researchers at The Scripps Research Institute have steadily built a model of how a powerful DNA repair complex works. Now, their latest discovery provides revolutionary insights into the way the molecular motor inside the complex functions – findings they say may have implications for treatment of disorders ranging from cancer to cystic fibrosis.
In a paper published in an Advance Online Edition of Nature Structural and Molecular Biology March 27, 2011, the scientists say that the complex's motor molecule, known as Rad50, is a surprisingly flexible protein that can change shape and even rotate depending on the task at hand.
The finding solves the long-standing mystery of how a single protein complex known as MRN (Mre11-Rad50-Nbs1) can repair DNA in a number of different, and tricky, ways that seem impossible for "standard issue" proteins to do, say team leaders Scripps Research Professor John Tainer, Ph.D., and Scripps Research Professor Paul Russell, Ph.D., who also collaborated with members of the Lawrence Berkeley National Laboratory on the study.
They say the finding also provides a critical insight into the ABC-ATPase superfamily of molecular motors, of which Rad50 is a member.
"Rad50 and its brethren proteins in this superfamily are biology's general motors," said Tainer, "and if we know how they work, we might be able to control biological outcomes when we need to."
For example, knowing that Rad50 changes its contour to perform a function suggests it might be possible to therapeutically target unique elements in that specific conformation. "There could be a new generation of drugs that are designed not against an active site, like most drugs now (an approach that can cause side effects, but against the shape the protein needs to be in to work," Tainer said.
Russell added, "Proteins are often viewed as static, but we are showing the moving parts in this complex. They are dynamic. They move about and change shape when engaging with other molecules."
First Responder
The MRN complex is known as a first-responder molecule that rushes in to repair serious double-strand breaks in the DNA helix—an event that normally occurs about 10 times a day per cell due to ultraviolet light and radiation damage, etc. If these breaks are not fixed, dangerous chromosomal rearrangements can occur that lead to cancer. Paradoxically, the complex also mends DNA breaks promoted by chemotherapy, protecting cells against cancer treatment.
When MRN senses a break, it activates an alarm telling the cell to shut down division until repairs are made. Then, it binds to ATP (an energy source) and repairs DNA in three different ways, depending on whether two ends of strands need to be joined together or if DNA sequences need to be replicated. "The same complex has to decide the extent of damage and be able to do multiple things," Tainer said. "The mystery was how it can do it all."
To find out, Tainer, head of a structural biology group, and Russell, who leads a yeast genetics laboratory, began collaborating five years ago. With the additional help of team members at Lawrence Berkeley National Laboratory and its Advanced Light Source beamline, called SIBYLS, the collaboration has produced a series of high-resolution images of the crystal structure of parts of all three proteins (rad50, Mre11, and Nbs1), taken from fission yeast and archaea. The scientists also used the lab's X-ray scattering tool to determine the proteins' overall architecture in solution, which approximates how a protein appears in a natural state.
The scientists say that the parts of the complex, when imagined together as a whole unit, resemble an octopus: the head consists of the repair machinery (the Rad50 motor and the Mre11 protein, which is an enzyme that can break bonds between nucleic acids) and the octopus arms are made up of Nbs1 which can grab the molecules needed to help the machinery mend the strands.
In this study, Tainer and Russell were able to produce crystal and X-ray scattering images of parts of where Rad50 and Mre11 touched each other, and what happened when ATP bound to this complex and what it looked like when it didn't.
In these four new structures, they showed that ATP binding allows Rad50 to drastically change its shape. When not bound to ATP, Rad50 is flexible and floppy, but bound to ATP, Rad50 snaps into a ring that presumably closes around DNA in order to repair it.
"We saw a lot of big movement on a molecular scale," said Tainer. "Rad50 is like a rope that can pull. It appears to be a dynamic system of communicating with other molecules, and so we can now see how flexibly linked proteins can alter their physical states to control outcomes in biology."
"We thought ATP allowed Rad50 to change shape, but now we have proof of it and how it works," Russell said. "This is a key part of the MRN puzzle."
An Engine for Many Vehicles
Rad50 and ATP provide the motor and gas for a number of biological machines that operate across species. These machines are linked to a number of disorders, such as cystic fibrosis, which is caused by a defect in the cystic fibrosis transmembrane conductance regulator (CFTR) gene, which is a member of the ABC ATPase superfamily.
"Our study suggests that ABC ATPase proteins are used so often in biology because they can flexibly hook up to so many different things and produce a specific biological outcome," Tainer said.
Given this new prototypic understanding of these motors, Tainer and Russell envision a future in which therapies might be designed that target Rad50 when it changes into a shape that promotes a disease. For example, chemotherapy could be coupled with an agent that prevents the MRN complex from repairing DNA damage, promoting death of cancer cells.
"There are some potentially very cool applications to these findings that we are only beginning to think about," Russell said.
Co-authors of the paper, "ABC ATPase signature helices in Rad50 link nucleotide state to Mre11 interface for DNA repair," include Gareth J. Williams, Soumita SilDas, and Michal Hammel of the Lawrence Berkeley National Laboratory; and R. Scott Williams, Jessica S. Williams, Gabriel Moncalian, Andy Arval, Oliver Limbo, and Grant Guenther of The Scripps Research Institute. See http://www.nature.com/nsmb/journal/vaop/ncurrent/abs/nsmb.2038.html.
The study was funded by the National Cancer Institute, the National Institutes of Health, and the Department of Energy.
Send comments to: mikaono[at]scripps.edu
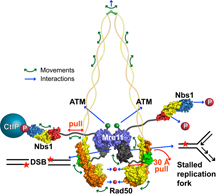
The new finding solves the long-standing mystery of how a single protein complex known as MRN (shown here) can repair DNA in a number of different ways. (Click to enlarge.)
