

Team Finds Immune Molecule that Attacks Wide Range of Flu Viruses
By Renee Twombly and Mika Ono
Researchers at The Scripps Research Institute report the characterization of an immune system molecule that targets what appears to be an "Achilles heel" of a wide range of influenza viruses – including the viruses responsible for past global pandemics, those causing current common infections, and strains of bird flu believed to pose future world threats.
The discovery of the molecule, an antibody known as CR6261, is good news for researchers who hope to design a flu vaccine that would give humans lifelong protection against a majority of influenza viruses. The antibody also has the potential to treat those who are unvaccinated and become infected with the flu.
The team's findings were published in the February 26, 2009, issue of Science Express, an advance, online publication of selected research papers from the prestigious journal Science.
"This is very exciting because it marks the first step toward the Holy Grail of influenza vaccinology – the development of a durable and cross-protective universal influenza virus vaccine," says the study's senior investigator, Ian Wilson, a professor in the Department of Molecular Biology and a member of The Skaggs Institute for Chemical Biology at Scripps Research. "Such a flu vaccine could be given to a person just once and act as a universal protectant for most subtypes of influenza, even against pandemic viruses."
Flu vaccines now offer protection only for the specific strains of influenza that public health officials believe to be currently circulating in the population. This involves a lot of guesswork about which strains will be most prevalent and, because the virus is constantly mutating, this guesswork must be repeated year after year.
According to the U.S. Centers for Disease Control, in the United States more than 200,000 people are typically hospitalized from flu complications every year, and about 36,000 people die from the illness. But that is in a normal year. Over the past century, three major human influenza pandemics (the Spanish Flu of 1918-1919, the Asian Flu of 1957-1958, and the Hong Kong Flu of 1968-1969) have devastated the human population, killing around 50-100 million people worldwide.
Broad Action
In the new research paper, the scientists, composed of a team from Scripps Research and the biopharmaceutical company Crucell, in the Netherlands, show that the CR6261 antibody attaches to the virus that caused the devastating 1918 "Spanish flu" and to a virus of the "H5" class of avian influenza that jumped from chickens to a human in Vietnam in 2004 The scientists at Crucell previously demonstrated in laboratory experiments that this antibody can neutralize common, seasonal flu viruses.
"We can see exactly how and where the antibody grabs on to these influenza viruses," says the study's first author, Damian Ekiert, a graduate student in the Scripps Research Kellogg School of Science and Technology working in the Wilson laboratory. "And we can see that this same mode of interaction occurs in viruses that are very different from each other."
Wilson says the discovery was possible because of the modern tools that the research team employed, such as phage display to isolate antibodies from human blood, and a state-of-the-art robotic crystallization laboratory that helps solve the structures of microbial antigens much more quickly than in the past.
"I have been working with influenza virus antigens since 1987, and I find it just amazing to suddenly see antibodies now appear that we had no idea existed," Wilson says.
Researchers at Scripps Research and collaborating institutions have long been looking for influenza antibodies with a broader spectrum of action. To find these antibodies, the researchers extracted white blood cells from a healthy immunized volunteer to make a library of antibodies to look for antibodies that interact with viruses that the donors could not have come into contact with before, such as H5 avian influenza that has spread only from chickens to humans, but not from humans to humans.
The researchers found one such antibody in the blood of a donor who had recently been vaccinated with a flu shot to protect against H1 influenza virus, one of the seasonal subtypes that most commonly circulates in humans. That antibody was isolated and named CR6261—although some of the researchers later dubbed it "Supermantibody" when they began to realize how effective it was.
CR6261-like antibodies have now also been found in other people. According to Ekiert, it is likely that many people, if not all, have these antibodies, but the body doesn't always produce or use them efficiently.
Solving the Puzzle
The next step for the researchers was to understand exactly how CR6261 recognized and responded to such a broad array of influenza viruses.
To do that, Ekiert led the successful effort to solve two crystal structures: one with the antibody bound to the hemagglutinin H1 virus that caused the 1918 pandemic and another with the antibody glued to the hemagglutinin from the 2004 Vietnam H5 avian influenza.
Influenza antibodies, including those induced by current vaccines, target mushroom-shaped proteins known as hemagglutinin (HA) that stud the outer coat of a virus particle to help the virus infect cells of a host organism, such as humans.
What the Scripps Research scientists found is that CR6261 latches on to the "stalk" of the mushroom-like hemagglutinin particle, near where the protein juts out from the viral coat, and that this binding area, known as an epitope, is the same in both the H1 and H5 viruses. The scientists then analyzed the genome of more than 5,000 different influenza viruses and found the epitope's sequence is nearly identical in all of them, suggesting that this part of the virus is much more highly conserved than the virus's constantly mutating cap.
This insight into the way the CR6261 antibody binds to the virus's structure makes sense, the researchers say. It helps explains why the antibody may not be as powerful as it needs to be to attack influenza. "The epitope it needs to latch on to is at the base of the stalk of the hemagglutinin protein, so it is difficult to get to because these proteins are packed together tightly on the viral coat," Ekiert says. "Plus, most antibodies try to attack the mushroom cap of the hemagglutinin proteins because that is much more accessible, and so this probably sets up a huge competition between antibodies."
"Certain regions of the hemagglutinin protein are like big red flags to the immune system, but they are functionally unimportant," Wilson says. "The task now is to figure out how to suppress reactivity with those regions and enhance the immune system's attack on this conserved epitope."
It may also be possible that some people who rarely if ever contract the flu may have CR6261-like antibodies that are more efficient than others in neutralizing influenza viruses.
So far, the researchers have shown the CR6261 antibody works against many of the 16 different subtypes of influenza viruses. The antibody neutralized every H1 virus that the group tested, including those that have caused pandemics over the past 100 years. The antibody also worked on the H5 bird viruses that are not yet circulating in humans. However, the CR6261 antibody is not effective for the H3 subclass, which is a common human influenza virus, because a sugar molecule blocks the epitope.
"If a sugar is the only impediment in the way, we think there is a way around that in vaccine design," Wilson says. "Even so, this antibody could still potentially hit 12 out of the 16 influenza viral subtypes. We now have a blueprint upon which to design the next generation of anti-virals, and that is why we are so enthusiastic about these findings as they give hope that it may indeed be possible to generate a universal vaccine against influenza virus, as well as provide immediate protection when used as an antibody therapeutic."
In addition to Wilson and Ekiert, authors of the paper "Antibody recognition of a highly conserved epitope across influenza viruses" are Gira Bhabha and Marc-André Elsliger of Scripps Research, and Robert Friesen, Mandy Jongeneelen, Mark Throsby, and Jaap Goudsmit of Crucell Holland BV, Leiden, The Netherlands.
The work was funded by a grant from the National Institutes of Health, a predoctoral fellowship from the ARCS Foundation and the Skaggs Institute. Facilities supporting this work were funded by the NIH National Institute of General Medical Sciences, the National Cancer Institute, and the U.S. Department of Energy.
Send comments to: mikaono[at]scripps.edu

"This is very exciting because it marks the first step toward the Holy Grail of influenza vaccinology—the development of a durable and cross-protective universal influenza virus vaccine," says Professor Ian Wilson.

"We can see exactly how and where the antibody grabs on to these influenza viruses," says the study's first author, graduate student Damian Ekiert.
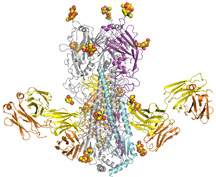
An image of the antibody
known as CR6261, bound to the hemagglutinin spike of the influenza virus
that caused a lethal human case of bird flu in 2004. Click to enlarge
