

Scientists Reveal Key Structure from Ebola Virus
By Jason Socrates Bardi
Scientists at The Scripps Research Institute have determined the structure of a critical protein from the Ebola virus, which, though rare, is one of the deadliest viruses on the planet killing between 50 and 90 percent of those infected.
Described in the cover article of the July 10, 2008 issue of the journal Nature, the research reveals the shape of the Ebola virus spike protein, which is necessary for viral entry into human cells, bound to an immune system antibody acting to neutralize the virus. The structure provides a major step forward in understanding how the deadly virus works, and may be useful in the development of potential Ebola virus vaccines, or treatments for those infected.
"Much about Ebola virus is still a mystery," says Erica Ollmann Saphire, the Scripps Research scientist who led the five-year effort. "However, this structure now reveals how this critical piece of the virus is assembled and, importantly, identifies vulnerable sites that we can exploit."
There is currently no cure for Ebola hemorrhagic fever. The virus is spread when people come into contact with the bodily fluids of someone who is already infected. Most ultimately die from a combination of dehydration, massive bleeding, and shock. The best treatment consists of administering fluids and taking protective measures to ensure containment, like isolating the patient and washing sheets with bleach.
The breakthrough described in the Nature article, though, provides hope that one day modern medicine will have more to offer.
The structure of the antibody together with the viral glycoprotein helps reveal the mechanisms by which the molecules assemble on the viral surface and helps explain how the pathogen evades and exploits the human immune system. The structure also provides a guide for the design of drugs and vaccines to block this protein, potentially preventing disease and death.
The new research was made possible by an antibody isolated by Dennis Burton, a Scripps Research professor and one of the study's coauthors. The antibody—shown bound to the Ebola virus spike protein in the current research—was derived from bone marrow of one of the few survivors of the 1995 Ebola outbreak in Kikwit, a city in the southwestern part of the Democratic Republic of Congo. The Kikwit outbreak was particularly deadly, with a higher than 90 percent mortality rate for those infected.
In addition to its importance for Ebola, the new research has broader implications for the study of viruses in general.
"Structures of the native, oligomeric forms of viral glycoproteins as they exist on the viral surface are exceedingly difficult to achieve and thus exceedingly rare," notes Ollmann Saphire. "This structure now provides templates by which researchers studying other viruses could try to understand how their virus's surface protein is assembled and neutralized by an antibody."
In addition to Ollmann Saphire, the article, "Structure of the trimeric, prefusion Ebola virus glycoprotein in complex with a neutralizing antibody from a human survivor," was authored by Jeffrey E. Lee, Marnie L. Fusco, Ann J. Hessell, Wendelien B. Oswald, and Dennis R. Burton at The Scripps Research Institute. For more information on the paper, see the journal Nature for the paper's abstract (with links to the full text) at
http://www.nature.com/nature/journal/v454/n7201/abs/nature07082.html and the article "Making the Paper: Erica Ollmann Saphire" at
http://www.nature.com/nature/journal/v454/n7201/full/7201xa.html.
Support for the research was provided by grants from the National Institutes of Health, a Career Award from the Burroughs Wellcome Fund, and a fellowship from the Canadian Institutes of Health Research.
For more information about the Ollmann Saphire lab, including an animated rotating model of the new structure, see http://www.scripps.edu/ims/ollmannsaphire/research.htm.
Send comments to: mikaono[at]scripps.edu
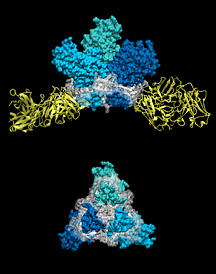
These images show different perspectives of the Ebola virus spike protein (blue and white), which is necessary for viral entry into human cells, bound to an immune system antibody (yellow) acting to neutralize the virus. Click to enlarge.
