

Scientists Develop Modeling Method that Increases Our Understanding of Diseases like Cystic Fibrosis and Alzheimer's
By Jason Bardi
A team of scientists at The Scripps Research Institute has come up with a simple but comprehensive way of probing the parts of our biological machinery that controls protein folding, packaging, and export from our cells.
The team has created a new model integrating the chemistry and biology of protein folding, called folding for export (FoldEx), which provides a general framework for understanding the causes and potential treatment strategies of diseases that arise when this protein homeostasis machinery malfunctions. These diseases include type 2 diabetes, Gaucher's disease, cystic fibrosis, and Alzheimer's disease.
The team describes its new mathematical model of the chemistry and biology of cellular protein homeostasis in the latest issue of the journal Cell, published on November 16.
"If we can begin to understand how the cellular folding, packaging, and export pathways work," says Jeffery Kelly, who is the Lita Annenberg Hazen Professor of Chemistry and a member of the The Skaggs Institute for Chemical Biology at Scripps Research, "this should give us insight into how to ameliorate diseases that arise when these pathways get out of balance."
"Our new model allows us to think about therapeutic intervention in a completely new way," says Professor William Balch, a Scripps Research cell biology professor and a member of the Scripps Research Institute of Childhood and Neglected Diseases.
Balch and Kelly guided the creation of the FoldEx model with Professor Joel Buxbaum. The new approach began as a back-of-the-envelope idea and was made a reality by the combined efforts of Assistant Professor Evan Powers and Luke Wiseman, a recent graduate of the Kellogg School of Science and Technology at Scripps Research.
Biological Machinery in Health and Disease
In many heritable diseases, specific mutations within a particular gene simply cause the protein product of the gene to malfunction. But often a defect in the general biological machinery of the cell that interacts with the protein is also involved, causing a loss of activity because of this machinery's crucial role in protein expression and function.
Gene expression starts when DNA is transcribed into RNA and is then translated into proteins, but the process does not end there. Proteins are unfolded when first expressed, and they usually must fold into a compact, three-dimensional structure in order to function properly. The biological machinery helps them achieve this by providing special molecules called chaperones and folding enzymes that help proteins fold.
Quite often, proteins also need help to get to where they are going. Some have to be transported and inserted into a cell's membrane. Others are intended to be secreted into the bloodstream or surrounding tissues, and have to be transported outside the cell. The biological machinery of the cell accomplishes all this starting with a tubular labyrinth called the endoplasmic reticulum (ER). The ER is like a massive shipping facility that folds, packages, and exports proteins to destinations inside and outside of the cell. About a third of the proteins in the human body go through the ER.
The biological machinery of the ER also protects the body by degrading potentially dangerous proteins. Sometimes it is too efficient at doing so, and many diseases can be traced to this action. Cystic fibrosis, for instance, is caused by the loss of an essential protein called cystic fibrosis transmembrane conductance regulator (CFTR), a chloride channel, that regulates hydration of the lung surface. Without CFTR, the mucous in the lungs becomes thick, sticky, and prone to harboring bacterial infections—the classic symptoms of cystic fibrosis.
People with cystic fibrosis have mutations in their CFTR protein, and the biological machinery responds to these mutations by targeting mutated CFTR for degradation. Thus, it is not that people with cystic fibrosis cannot make CFTR, but that the protein they do make simply can never make it to the cell surface. A single mutation in CFTR is enough to cause it to misfold, become degraded, and cause cystic fibrosis.
Many other diseases result from a similar malfunction, including Gaucher disease, which is caused by the accumulation of a fatty substance in the spleen, liver, lungs, bone marrow, and sometimes the brain because of the loss of the lipid metabolizing enzyme that breaks it down. Gaucher disease is often treated, in fact, by administering a recombinant form of the missing enzyme.
A converse problem occurs when the biological machinery exports proteins out of the ER that then misfold and cause destruction once they get to their destination. A number of rare and common conditions termed "amyloid" diseases stem from this etiology. Familial amyloid polyneuropathy (FAP), for instance, results from the misfolding and deposition of one of 100 mutants of the protein transthyretin. An analogous disease called familial amyloid cardiomyopathy (FAC) causes fibril formation in the heart and leads to cardiac dysfunction. About one million African-Americans carry the gene that predisposes them to FAC. Another amyloid disease affecting the heart, Senile Systemic Amyloidosis (SSA), afflicts an estimated 10 to 15 percent of all Americans over the age of 60.
Moreover, the deposition of protein plaques in the brain is seen in diseases like Alzheimer's, Parkinson's, and Huntington's, though it is not yet clear whether these plaques are the cause or a downstream effect of the disease.
For years, scientists have recognized that these diseases all in some way relate to the underlying biological machinery, which protects us when we are young. But how can scientists approach tinkering with this biology to treat these diseases? That is exactly the question that the FoldEx mathematical model addresses.
The Model and What It Predicts
FoldEx started with a conversation three years ago among Wiseman, Powers, and Buxbaum on how the biological machinery of the ER is related to misfolding diseases.
Soon the conversation grew to involve Kelly, Balch, and nearly everybody in their two laboratories. The product of this work, after many, many rounds of refinement, was a sophisticated analysis that treats rather complex competing pathways as if they were single enzymes, greatly simplifying the analysis without loosing the essence of the chemistry and biology of protein homeostasis.
Because of this simplicity, FoldEx can represent complicated processes with well-established biochemical principles that help explain how these pathways compete for various conformational ensembles of proteins and dictate the balance between folding, secretion, and degradation. But this simplification does not come at the expense of relevance. In the Cell paper, the Scripps Research team showed that FoldEx can predict concepts, principles, and results that have already been validated. The real value of the model, however, is that it can be used to predict how folding and misfolding diseases may be delayed or reversed by manipulating the innate biological machinery.
FoldEx shows that by adjusting the folding and export machinery, it may be possible to address misfolding diseases without replacing the problematic proteins. The model predicts how the system would respond if you tweak it here or there to normalize the physiology—for instance by increasing degradation to reduce export and therefore reduce the amount of amyloid protein that gets deposited in tissues like the brain.
Likewise, one can imagine adjusting the levels of chaperone helper molecules so that a fraction of the proteins that are normally degraded will be folded and exported to restore function in loss-of-function diseases. The idea is that if partially functional folded mutant proteins can be delivered to where they are needed, they will be active enough to avoid disease.
What this means, say Balch and Kelly, is that you aren't necessarily destined to get a disease even if you've inherited a bad mutation if we can learn to adjust the fold or the biological folding environment. Here, a severe mutation could be viewed as a more benign polymorphism.
The article in Cell is titled"An Adaptable Standard for Protein Export from the Endoplasmic Reticulum."
Support for this work was provided by the National Institutes of Health, the Skaggs Institute for Chemical Biology, the Lita Annenberg Hazen Foundation, the Cystic Fibrosis Foundation, a Norton B. Gilula Fellowship, and a Fletcher Jones Foundation Fellowship.
For Science Watch, at the end of the second last paragraph, add: See
http://www.cell.com/content/article/abstract?uid=PIIS0092867407013438.
Send comments to: mikaono[at]scripps.edu
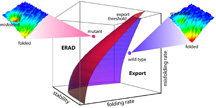
The new approach, called "folding for export" or "FoldEx," integrates the chemistry and biology of protein folding to provide a general framework for understanding the causes of—and potential treatment strategies for—a number of diseases. Click for image details.
