

Physicist in Disguise
By Eric Sauter
When Ruben Abagyan was 13-years-old, his father, a world-renowned Soviet nuclear physicist, told his son that the age of nuclear physics was over, that the scientific problems in the field had long been solved and that he should start looking elsewhere.
"He said to me, 'If you want to do exciting science, you have to apply physics to biology," Abagyan said, "and I followed his advice."
Nearly 40 years later, the old Soviet Union gone, Ruben Abagyan is still applying his knowledge of physics to the problems of biology and doing it successfully at The Scripps Research Institute, the place that has been his scientific home since 1999.
His laboratory works in the area of computational structural biology, one tributary among many of a much larger river called computational biology—which weaves together any number of disciplines (mathematics, physics, chemistry, computers, statistics, proteomics; the list goes on)—to attack biological problems on the molecular level using computer programs. For Abagyan, that means producing a brand of molecular prediction that goes, as he says, "beyond what you can approach with experimental methods."
Practical Results
Recently, Abagyan and several collaborators from places as diverse as Columbus, Ohio, and Melbourne, Australia, did just that, using the predictive power of computers to uncover a new and potentially powerful therapeutic for prostate cancer from a library of drugs already on the market—in this case, the phenothiazine antipsychotics. The process—called "drug repurposing"—was published in June by the Proceedings of the National Academy of Sciences.
"I'm a physicist in disguise," Abagyan said, "because I talk on some level about a lot of different things. But while the core methodology we use is physics—molecular physics, some aspects of chemical physics—we want to answer biological questions."
In Abagyan's latest study, his computational biology approach managed two unique feats at the same time. First, the study uncovered a secondary effect of a well-known drug, a process that can be repeated with different drugs and different drug targets. Second, it found a new chemical scaffold for androgen receptor antagonists, an important discovery in and of itself because, as he points out, there simply aren't enough of them around that can be used against prostate cancer.
All drugs have activity in addition to the interaction with their selected target; generally this gets shelved under the side effects profile. A significant portion of Abagyan's work was aimed specifically at these secondary targets, enhancing the secondary activity and eliminating the original target. The results, as the study notes, are drugs that "are likely to enter clinical trials more rapidly and at less cost," and with proven safety and bioavailability profiles.
"I wouldn't say our new study is the quintessential example [of computational biology], but it is a very interesting application of computational structural biology," he said. "Some people think the whole process is too theoretical, but our structural predictive approach brings a very practical result and replaces cumbersome, expensive, and error-prone high-throughput screening techniques. This is the elegant approach—it's not messy."
Abagyan begins with a receptor structure or its approximate 3D model and, using several advanced computer algorithms, predicts location, properties, and dynamics of the binding pocket. A large collection of those pockets, the pocketome, then enables ligand docking and virtual screening to detect inhibitors of specific molecular targets. Over the past several years, the lab's efforts in the last area have led to new or improved inhibitors of the thyroid hormone receptor, retinoic acid receptor, receptor for epidermal growth factor (EGFR), anthrax lethal factor, malarial enoyl carrier-protein reductase, dynamin, alpha-1-antitrypsin, and, finally, the androgen receptor . The list of "conquered" biomedical targets is growing fast.
"Crystallography alone gives only partial information about protein A1 in complex with ligand L," he said. "In reality, we want to predict what will happen between protein A2, or even its homologue B, and ligand M—and it is only with computational methods that we can instantly travel to reach another functional state, another receptor, and another ligand."
It was one of Abagyan's programs—ICM or Internal Coordinate Mechanics—that solved many key problems in predicting structure and interactions.
Back in the USSR
Although he isn't entirely comfortable with the characterization, Abagyan is something of a walking encapsulation of modern scientific history; more specifically, the sudden rise to dominance of Soviet science and its equally sudden downfall.
Abagyan was born in 1957 in the town of Obninsk, just 100 kilometers southwest of Moscow. Obninsk, which in its prime housed "no fewer than 12 scientific research institutions and a technical university," according to Wired magazine, was the site of the world's first operational nuclear power plant. Abagyan's parents were both physicists and Abagyan himself studied molecular physics at the Moscow Institute of Physics and Technology, perhaps the most prestigious of all the Soviet-era research institutions.
Abagyan studied, worked, and got married (his wife, Margarita, was trained as a bio-physicist) in the Soviet Union but left in 1989 just before the collapse of the old regime. What he remembers of the country growing up is much different than the one he left.
"We—I mean my parents and our friends—were completely focused on physics and mathematics. In our backyard you could find professors, researchers, any number of outstanding scientists. This was the age of Sputnik and Yuri Gagarin, a time when Soviet science and technology was admired around the world. There was also a great sense that your life was going to be fine, that the state would take care of everything."
But 25 years and five general secretaries of the Communist Party later it came to an end.
Abagyan was 32 years old when he left, going first to Belgium and then to Heidelberg, Germany, were he settled in at the famous European Molecular Biology Laboratory. Given his background, he was already something of a known quantity, and he was the first scientist from the East offered a full staff position. He stayed for five years.
In 1994, he moved to New York University, and became the director of computational biology at the Skirball Institute of Biomolecular Medicine and tenured associate professor of biochemistry and mathematics. Soon afterwards, he and his wife and his children moved to New Jersey where they settled into suburban life. It was at Skirball that he developed his practical approach to science, in fact, to running a laboratory in general: "I like multi-tasking. I can prioritize and delegate. Once I find the right people all I need to do is spot check."
The Bottleneck Philosophy
It was also in New York that he began a company based around the program he had developed in Heidelberg.
The company, first named Biosoft and then renamed Molsoft in 1995, still thrives in La Jolla where the couple and his main research partner, MolSoft Principal Scientist Maxim Totrov, moved in 1999. The ICM program has been in constant evolution ever since. The basic algorithms used in ICM have made it an effective platform for everything from peptide prediction to flexible macromolecular docking. As the biological problems have grown more complex, the program has expanded into new areas, such as graphics, chemistry, database searches, and statistics.
"I have these five-year plans," he said with obvious irony. "In 1999, I got a call from Peter Schultz (the Schultz lab at Scripps Research also works in the area of molecular structure with an eye on creating new molecules) who was just starting up the Novartis Institute, so I became one of the first people to join."
He soon left Novartis to join Scripps Research.
And, as was the case when he started Molsoft, his main concern was maintaining the ability to develop the program as he saw fit.
"The practical direction is one of the directions I like to take with our research," he said. "It is the basic philosophy that unifies all our different projects—we've had recent papers that are far from structural studies, things like chip design, or data dissemination, for example. I like to work in different areas."
Abagyan calls it the bottleneck philosophy.
"First, you identify an important practical application; and by that I mean something that affects the well being of many people. The suitable application has to have a problem or bottleneck in an area that can be attacked through our methods. So, we identify the bottleneck, and apply our protocol to see if we can solve it. Then you apply the improved method and move onto the next bottleneck."
The point being is to keep moving forward. "It keeps you humble," Abagyan said, "especially if you discover that the bottleneck isn't a something you're trained to solve. But you don't shy away from it. You go in and do what you have to do to solve it."
It also keeps things forever new: "Its like the old saying, you never step in the same river twice."
Send comments to: mikaono[at]scripps.edu

Professor Ruben Abagyan works in the area of computational structural biology, which he believes can go "beyond what you can approach with experimental methods. Photo by Kevin Fung.
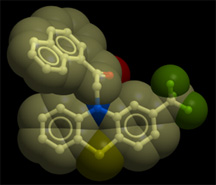
Using the predictive power of computers, the Abagyan lab has uncovered a new and potentially powerful therapeutic for prostate cancer from a library of drugs already on the market, the phenothiazine antipsychotics. This image shows the novel non-steroidal androgen receptor antagonist that resulted from "re-purposing" these drugs.
