

Study Shows How Genetic Repair Mechanism Helps Seal DNA Breaks
A new study by researchers from The Scripps Research Institute, Lawrence Berkeley National Laboratory, the Washington University School of Medicine, and the University of Maryland has provided a clearer picture of the final steps of a critical DNA repair process. When these repair processes go awry, cells can malfunction, die, or become cancerous.
The study was published in the October 20, 2006 issue of the journal Molecular Cell.
"These results are exciting because they reveal for the first time how these proteins can dynamically assemble and change their shape to join DNA ends during DNA replication and repair," said a senior author of the paper John Tainer, who is a professor at Scripps Research, member of Scripps Research's Skaggs Institute for Chemical Biology, and co-principal investigator of the Structural Cell Biology of DNA Repair project in Berkeley Lab's Life Sciences Division.
As the genetic material, DNA is surprisingly reactive and under continuous assault from environmental toxins and reactive cellular metabolites, so a means of repairing DNA damage is essential to maintaining the integrity of our genetic blueprint for future generations.
DNA ligases are enzymes that are an essential part of this process, repairing millions of DNA breaks generated during the normal course of a cell's lifetime. Because the reaction joining the ends of DNA strands to restore the double helix is catalyzed by ligase enzymes and because this reaction is essential and abundant in dividing cells, DNA ligases are attractive targets in the development of new treatments for cancer and other diseases.
Ligase does its job in concert with another ring-shaped protein known as a sliding clamp. Sliding clamps like the human PCNA protein are master regulators of DNA repair, providing docking sites that recruit repair enzymes to the site of damage.
In the recent study, the scientists applied several state-of-the-art techniques to visualize DNA ligase alone and in complex with PCNA, using proteins from a model organism called Sulfolobus solfataricus that has many of the same biochemical characteristics of multicelled organisms, including humans. To visualize these complex and dynamic structures at high resolution, the team used a combination of x-ray crystallography and small angle x-ray scattering (SAXS) at the SIBYLS beamline at Berkeley Lab's Advanced Light Source.
"This paper shows that the SIBYLS beamline is well suited to define dynamic interactions that control cell biology and processes such as cancer," said Tainer. "These reversible complexes are also critical to efforts in understanding and controlling microbial responses and pathways."
Prior to the experiment, the scientists expected that DNA ligase would curl up in complex with the ring-shaped PCNA protein. However, results showed that ligase remains in an open conformation enabling other repair proteins to bind PCNA until the DNA is engaged and ligase snaps shut. The closed conformation of DNA ligase bound to DNA was imaged in a separate study previously reported by the same group of investigators.
"Our [new] study shows that DNA ligase switches from an open, extended shape to a closed, circular shape as it joins together DNA strands," said Tom Ellenberger, DVM, Ph.D., a senior author of the paper and the Raymond H. Wittcoff Professor and head of the Department of Biochemistry and Molecular Biophysics at Washington University School of Medicine in St. Louis. "The ligase resembles a wristwatch that cinches around the DNA ends that are being joined together. When ligase stacks against PCNA and encircles the DNA, we think this interaction ejects other repair proteins from PCNA. In this role, ligase may serve as the final arbiter of DNA repair, certifying that the DNA is in pristine condition and ready for the final step of DNA end joining."
The challenge for the future will be to study the molecular choreography of ligase, PCNA, and DNA in the same experiment, which will require new methods of analyzing SAXS data.
In addition to Tainer and Ellenberger, authors of the study, "A Flexible Interface between DNA Ligase and PCNA Supports Conformational Switching and Efficient Ligation of DNA," are John M. Pascal, Oleg V. Tsodikov, Greg L. Hura, Wei Song, Elizabeth A. Cotner, Scott Classen, and Alan E. Tomkinson.
Funding from the National Cancer Institute, the National Institute of General Medical Sciences, and the U.S. Department of Energy supported this research.
Send comments to: mikaono[at]scripps.edu
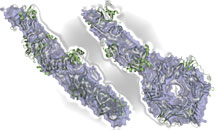
A new view of DNA ligase. Click here for details
