

Finger Exercises
By Eric Sauter
A pair of scientists from The Scripps Research Institute and their new web site were featured in a recent article describing a flurry of activity surrounding an engineered protein called a zinc finger. That's a term that doesn't come up much now in casual conversation but might in the future if estimates of its potential in gene therapy hold true. For gene therapy, the zinc finger nuclease binds to a specific mutated gene, cuts the DNA, and then repairs it with a non-mutated gene, hopefully eliminating the genetic disease in the process. The article, "Putting the Fingers on Gene Repair," appeared in the December 23, 2005 issue of the journal Science.
The article suggests that the Scripps Research laboratory of Professor Carlos Barbas may have hit on "…the simplest way to create a nuclease that targets enough DNA sequence to hit a specific gene…(through) the modular design approach" by stringing together a trio of different zinc fingers with a nuclease to target a specific DNA sequence. Barbas holds the Janet and Keith Kellogg II Chair in Molecular Biology and Chemistry and is a member of Scripps Research's Skaggs Institute for Chemical Biology. In an act of academic generosity, Jeff Mandell of the Barbas lab created a free web site called Zinc Finger Tools. On the site, which has been up and running since last September, biologists can type in a gene's DNA sequence and quickly learn how to create the right zinc fingers to target it.
The Zinc Finger Tools web site is to gene researchers what Home Depot is to home restoration fans—simple, utilitarian, and without a lot of frills. Underneath its plain vanilla exterior, the site is an ingenious search engine of enormous sophistication and breadth that saves valuable research time. Zinc Finger Tools can identify DNA targets for zinc finger proteins in just a few seconds, making it easy to scan any DNA sequence. The web site has already gone into Version 2.0, which fully enables the more complicated feature of designing zinc finger nucleases.
The idea for the web site came from the many requests the researchers received for zinc finger collaborations. "A lot of other labs approached us about it," says Research Associate Jeff Mandell. Mandell, who worked as a programmer at the San Diego Supercomputer Center before attending graduate school in biomedical sciences, joined the Barbas lab in November 2004. "Carlos thought it was a process that we could automate, so he asked me to scan some DNA sequences for target regions. It's difficult to design the finger transcription factors and there are a lot of little tricks to get it right. I realized you could put a number of tools on the site to make it easier—a kind of one-stop shopping for zinc finger researchers."
Zinc fingers are perhaps the ideal structure to build sequence-specific protein and Mandell's toolbox is filled to the brim with programs that can help do this. One program allows users to scan a given DNA sequence for consecutive DNA triplets that can be targeted with the zinc finger domains the Barbas lab has already published. Another tool—with the sparse but oddly seductive title "Design a Zinc Finger Protein"—uses a valid zinc finger target site, a DNA sequence for which the researchers have created novel zinc finger domains.
A third tool, "Predict Zinc Finger Protein Binding Site," allows subscribers to reverse engineer zinc finger proteins to determine the targeted DNA sequences, basically deconstructing the amino acid sequence of a zinc finger into its primary helices. If you know only the amino acid sequence of a zinc finger protein, as opposed to its DNA sequence, you can figure out the DNA sequence you need. A Mandell-designed algorithm searches for helices developed by the Barbas lab.
Registration for the site, which has received nearly 4,000 hits from close to 100 registered users, is quick and easy. "We don't review the registration, lock anybody out, or keep track of the information submitted to the site," Mandell says. The vast majority of users are academics from around the globe—most come from Europe, but Korea and Japan are represented as well.
The site's success, Mandell says, has been something of a surprise since they haven't publicized it much: "I think we sent the announcement to just a few people—so most of it has come from word of mouth."
The true value of the new web site, according to Barbas, is its sheer convenience.
"Currently, access to information on zinc finger DNA binding protein technology is not straightforward," he says. "While there are a number of reports claiming to make these proteins using novel but generally complex methods—for which validation is generally lacking—selecting one of these routes would mean spending more than a year to develop a protein sequence that our web site can generate in seconds. Our overall approach was not only the first to show that proteins could be custom-made to target specific DNA sequences and regulate genes, but also it's the only one that has been validated against multiple genes, in multiple species, and shown to work in whole organisms."
Barbas echoes Mandell's enthusiasm and predicts a great future for their effort: "Our web-based technology offers researchers an approach that allows them to reach into genomes to specifically regulate genes and to modify the hardware of genomes themselves. This is just the beginning. Even more high powered applications are on the horizon."
Send comments to: mikaono[at]scripps.edu
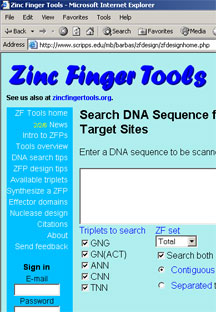
The Barbas lab has created a free web site for researchers called Zinc Finger Tools.
