
Xenopus tectum

Human iPSC-derived neurons

In vivo imaging of tectal NPCs and neurons

Phagocytic human iPSC-derived microglia
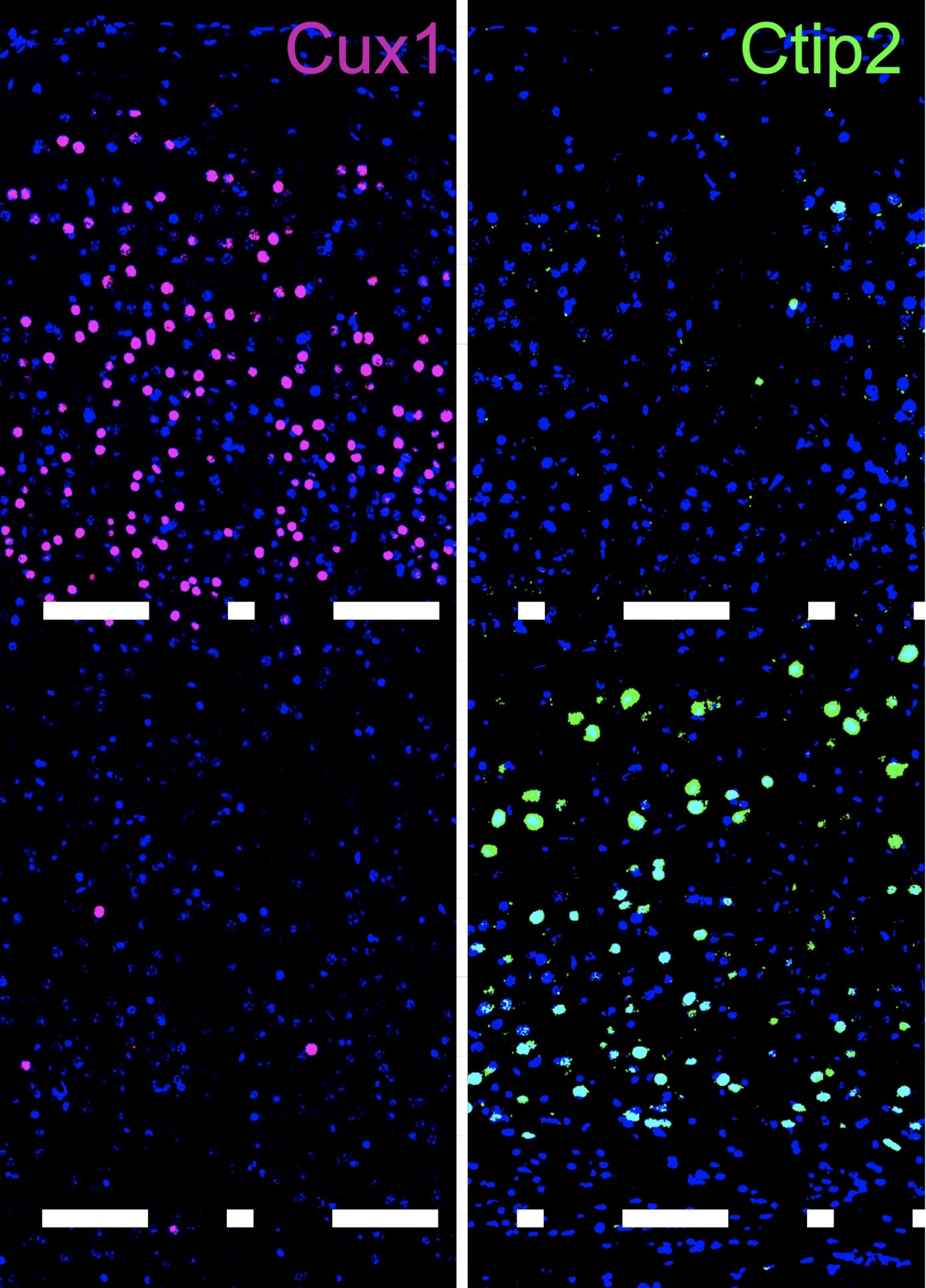
Mouse visual cortex lamination

Cell types in mouse brain
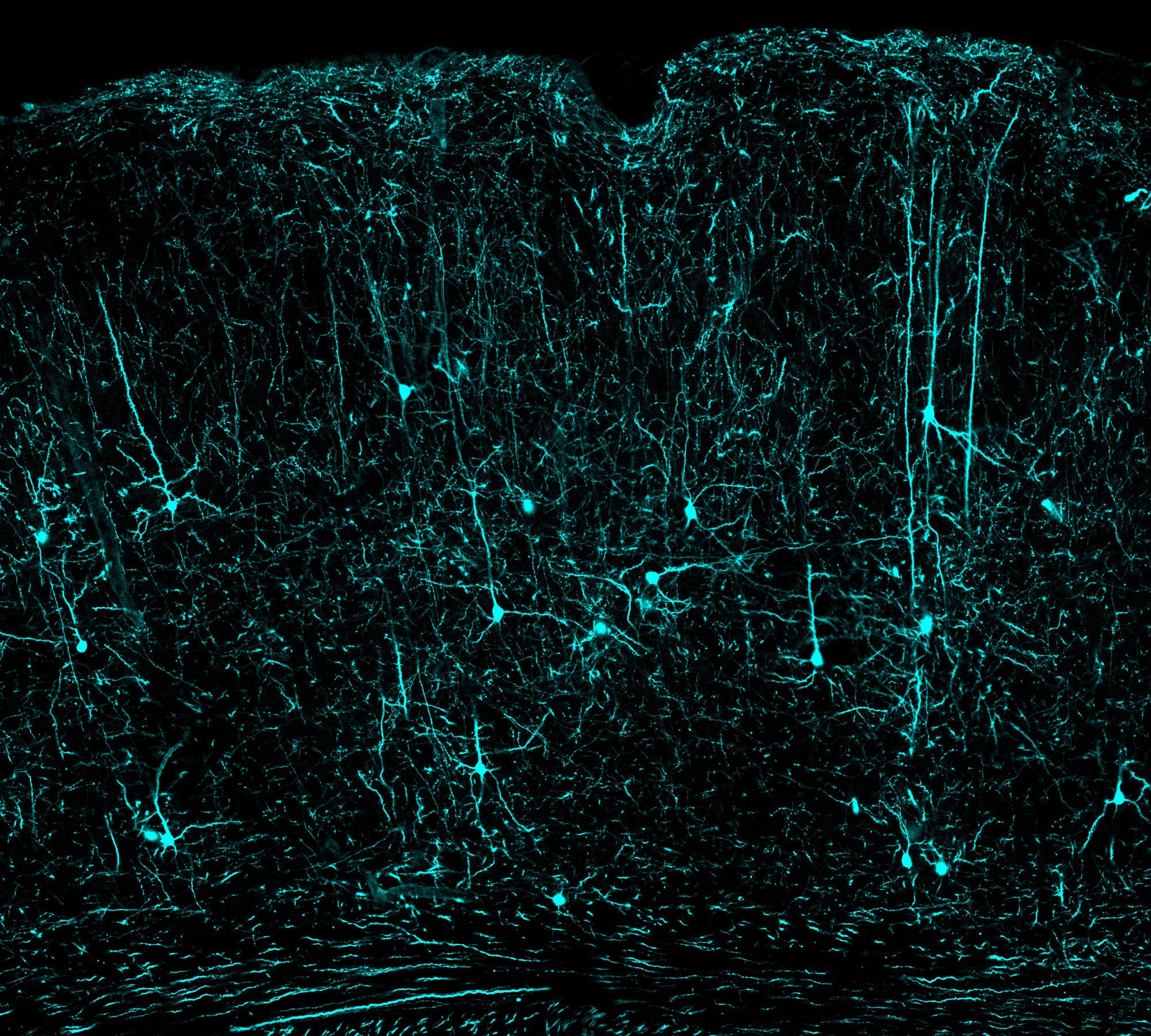
Mouse visual cortex
Studying the Role of Visual Experience and Intercellular Communication on Synaptic Dynamics
Our brains change over time as we develop, learn, and age. The Cline lab is interested in understanding the roles of experience and brain activity in orchestrating dynamic changes in the brain during development and aging. We are particularly interested in the role of visual experience in synaptic plasticity and recovery from injury of visual circuitry. We are also interested in extracellular vesicles as an avenue of intercellular communication that may have important roles in synaptic plasticity and neurological diseases.
